[English] 日本語
 Yorodumi
Yorodumi- EMDB-27078: Cryo-EM structures and computational analysis for enhanced potenc... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structures and computational analysis for enhanced potency in MTA-synergic inhibition of human protein arginine methyltransferase 5 | |||||||||
 Map data Map data | human protein arginine methyltransferase 5 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | PRMT5 / protein arginine methyl transferase / MTA-inhibitor synergy / cryo-EM structure-based drug design / computational analysis / catalytic mechanism / drug discovery / docking analysis / ONCOPROTEIN / ONCOPROTEIN-Transferase complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of adenylate cyclase-inhibiting dopamine receptor signaling pathway / peptidyl-arginine N-methylation / type II protein arginine methyltransferase / protein-arginine omega-N symmetric methyltransferase activity / peptidyl-arginine methylation / Golgi ribbon formation / negative regulation of epithelial cell proliferation involved in prostate gland development / oocyte axis specification / secretory columnal luminar epithelial cell differentiation involved in prostate glandular acinus development / histone H4R3 methyltransferase activity ...positive regulation of adenylate cyclase-inhibiting dopamine receptor signaling pathway / peptidyl-arginine N-methylation / type II protein arginine methyltransferase / protein-arginine omega-N symmetric methyltransferase activity / peptidyl-arginine methylation / Golgi ribbon formation / negative regulation of epithelial cell proliferation involved in prostate gland development / oocyte axis specification / secretory columnal luminar epithelial cell differentiation involved in prostate glandular acinus development / histone H4R3 methyltransferase activity / epithelial cell proliferation involved in prostate gland development / protein-arginine N-methyltransferase activity / methylosome / positive regulation of mRNA splicing, via spliceosome / methyl-CpG binding / endothelial cell activation / histone H3 methyltransferase activity / histone methyltransferase activity / regulation of mitotic nuclear division / negative regulation of gene expression via chromosomal CpG island methylation / Cul4B-RING E3 ubiquitin ligase complex / histone methyltransferase complex / E-box binding / positive regulation of oligodendrocyte differentiation / negative regulation of cell differentiation / spliceosomal snRNP assembly / ribonucleoprotein complex binding / regulation of ERK1 and ERK2 cascade / ubiquitin-like ligase-substrate adaptor activity / liver regeneration / regulation of signal transduction by p53 class mediator / methyltransferase activity / circadian regulation of gene expression / DNA-templated transcription termination / Regulation of TP53 Activity through Methylation / RMTs methylate histone arginines / protein polyubiquitination / p53 binding / transcription corepressor activity / snRNP Assembly / ubiquitin-dependent protein catabolic process / transcription coactivator activity / chromatin remodeling / protein heterodimerization activity / positive regulation of cell population proliferation / regulation of DNA-templated transcription / regulation of transcription by RNA polymerase II / chromatin / Golgi apparatus / nucleoplasm / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.14 Å | |||||||||
 Authors Authors | Yadav GP / Wei Z / Xiaozhi Y / Chenglong L / Jiang Q | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Commun Biol / Year: 2022 Journal: Commun Biol / Year: 2022Title: Cryo-EM structure-based selection of computed ligand poses enables design of MTA-synergic PRMT5 inhibitors of better potency. Authors: Wei Zhou / Gaya P Yadav / Xiaozhi Yang / Feng Qin / Chenglong Li / Qiu-Xing Jiang /  Abstract: Projected potential of 2.5-4.0 Å cryo-EM structures for structure-based drug design is not well realized yet. Here we show that a 3.1 Å structure of PRMT5 is suitable for selecting computed ...Projected potential of 2.5-4.0 Å cryo-EM structures for structure-based drug design is not well realized yet. Here we show that a 3.1 Å structure of PRMT5 is suitable for selecting computed poses of a chemical inhibitor and its analogs for enhanced potency. PRMT5, an oncogenic target for various cancer types, has many inhibitors manifesting little cooperativity with MTA, a co-factor analog accumulated in MTAP-/- cells. To achieve MTA-synergic inhibition, a pharmacophore from virtual screen leads to a specific inhibitor (11-2 F). Cryo-EM structures of 11-2 F / MTA-bound human PRMT5/MEP50 complex and its apo form resolved at 3.1 and 3.2 Å respectively show that 11-2 F in the catalytic pocket shifts the cofactor-binding pocket away by ~2.0 Å, contributing to positive cooperativity. Computational analysis predicts subtype specificity of 11-2 F among PRMTs. Structural analysis of ligands in the binding pockets is performed to compare poses of 11-2 F and its redesigned analogs and identifies three new analogs predicted to have significantly better potency. One of them, after synthesis, is ~4 fold more efficient in inhibiting PRMT5 catalysis than 11-2 F, with strong MTA-synergy. These data suggest the feasibility of employing near-atomic resolution cryo-EM structures and computational analysis of ligand poses for small molecule therapeutics. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27078.map.gz emd_27078.map.gz | 59.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27078-v30.xml emd-27078-v30.xml emd-27078.xml emd-27078.xml | 19.6 KB 19.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_27078.png emd_27078.png | 267.1 KB | ||
| Filedesc metadata |  emd-27078.cif.gz emd-27078.cif.gz | 6.6 KB | ||
| Others |  emd_27078_half_map_1.map.gz emd_27078_half_map_1.map.gz emd_27078_half_map_2.map.gz emd_27078_half_map_2.map.gz | 12.8 MB 12.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27078 http://ftp.pdbj.org/pub/emdb/structures/EMD-27078 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27078 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27078 | HTTPS FTP |
-Related structure data
| Related structure data |  8cyiMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27078.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27078.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | human protein arginine methyltransferase 5 | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.11 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half Map 1
| File | emd_27078_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half Map 2
| File | emd_27078_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of PRMT5 and MEP50 in the presence of MTA and a novel inh...
| Entire | Name: Complex of PRMT5 and MEP50 in the presence of MTA and a novel inhibitor (11-2F) |
|---|---|
| Components |
|
-Supramolecule #1: Complex of PRMT5 and MEP50 in the presence of MTA and a novel inh...
| Supramolecule | Name: Complex of PRMT5 and MEP50 in the presence of MTA and a novel inhibitor (11-2F) type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 440 KDa |
-Macromolecule #1: Protein arginine N-methyltransferase 5
| Macromolecule | Name: Protein arginine N-methyltransferase 5 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: type II protein arginine methyltransferase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 72.766664 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAAMAVGGAG GSRVSSGRDL NCVPEIADTL GAVAKQGFDF LCMPVFHPRF KREFIQEPAK NRPGPQTRSD LLLSGRDWNT LIVGKLSPW IRPDSKVEKI RRNSEAAMLQ ELNFGAYLGL PAFLLPLNQE DNTNLARVLT NHIHTGHHSS MFWMRVPLVA P EDLRDDII ...String: MAAMAVGGAG GSRVSSGRDL NCVPEIADTL GAVAKQGFDF LCMPVFHPRF KREFIQEPAK NRPGPQTRSD LLLSGRDWNT LIVGKLSPW IRPDSKVEKI RRNSEAAMLQ ELNFGAYLGL PAFLLPLNQE DNTNLARVLT NHIHTGHHSS MFWMRVPLVA P EDLRDDII ENAPTTHTEE YSGEEKTWMW WHNFRTLCDY SKRIAVALEI GADLPSNHVI DRWLGEPIKA AILPTSIFLT NK KGFPVLS KMHQRLIFRL LKLEVQFIIT GTNHHSEKEF CSYLQYLEYL SQNRPPPNAY ELFAKGYEDY LQSPLQPLMD NLE SQTYEV FEKDPIKYSQ YQQAIYKCLL DRVPEEEKDT NVQVLMVLGA GRGPLVNASL RAAKQADRRI KLYAVEKNPN AVVT LENWQ FEEWGSQVTV VSSDMREWVA PEKADIIVSE LLGSFADNEL SPECLDGAQH FLKDDGVSIP GEYTSFLAPI SSSKL YNEV RACREKDRDP EAQFEMPYVV RLHNFHQLSA PQPCFTFSHP NRDPMIDNNR YCTLEFPVEV NTVLHGFAGY FETVLY QDI TLSIRPETHS PGMFSWFPIL FPIKQPITVR EGQTICVRFW RCSNSKKVWY EWAVTAPVCS AIHNPTGRSY TIGL UniProtKB: Protein arginine N-methyltransferase 5 |
-Macromolecule #2: Methylosome protein 50
| Macromolecule | Name: Methylosome protein 50 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 33.226211 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: PNAPACMERQ LEAARYRSDG ALLLGASSLS GRCWAGSLWL FKDPCAAPNE GFCSAGVQTE AGVADLTWVG ERGILVASDS GAVELWELD ENETLIVSKF CKYEHDDIVS TVSVLSSGTQ AVSGSKDICI KVWDLAQQVV LSSYRAHAAQ VTCVAASPHK D SVFLSCSE ...String: PNAPACMERQ LEAARYRSDG ALLLGASSLS GRCWAGSLWL FKDPCAAPNE GFCSAGVQTE AGVADLTWVG ERGILVASDS GAVELWELD ENETLIVSKF CKYEHDDIVS TVSVLSSGTQ AVSGSKDICI KVWDLAQQVV LSSYRAHAAQ VTCVAASPHK D SVFLSCSE DNRILLWDTR CPKPASQIGC SAPGYLPTSL AWHPQQSEVF VFGDENGTVS LVDTKSTSCV LSSAVHSQCV TG LVFSPHS VPFLASLSED CSLAVLDSSL SELFRSQAHR DFVRDATWSP LNHSLLTTVG WDHQVVHHVV PT UniProtKB: Methylosome protein WDR77 |
-Macromolecule #3: N-[(2-aminoquinolin-7-yl)methyl]-9-(2-hydroxyethyl)-2,3,4,9-tetra...
| Macromolecule | Name: N-[(2-aminoquinolin-7-yl)methyl]-9-(2-hydroxyethyl)-2,3,4,9-tetrahydro-1H-carbazole-6-carboxamide type: ligand / ID: 3 / Number of copies: 1 / Formula: P2R |
|---|---|
| Molecular weight | Theoretical: 414.5 Da |
| Chemical component information | 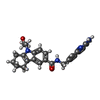 ChemComp-P2R: |
-Macromolecule #4: 5'-DEOXY-5'-METHYLTHIOADENOSINE
| Macromolecule | Name: 5'-DEOXY-5'-METHYLTHIOADENOSINE / type: ligand / ID: 4 / Number of copies: 1 / Formula: MTA |
|---|---|
| Molecular weight | Theoretical: 297.334 Da |
| Chemical component information | 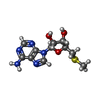 ChemComp-MTA: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.4 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 20 sec. / Pretreatment - Atmosphere: OTHER / Details: Preclean plasma cleaner |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK III |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Number real images: 4115 / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: OTHER / Imaging mode: DARK FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.5 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: OTHER / Overall B value: 68 |
|---|---|
| Output model |  PDB-8cyi: |
 Movie
Movie Controller
Controller



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)