+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Photosynthetic assembly of Chlorobaculum tepidum (RC-FMO2) | |||||||||
 Map data Map data | RC-FMO2 map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | FMO / reaction center / single-particle cryo-EM / bacteriochlorophyll / photosynthesis / electron transport chain / energy transfer | |||||||||
| Function / homology |  Function and homology information Function and homology informationthylakoid / bacteriochlorophyll binding / iron-sulfur cluster binding / photosynthesis / electron transfer activity / heme binding / membrane / metal ion binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Chlorobaculum tepidum TLS (bacteria) Chlorobaculum tepidum TLS (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.08 Å | |||||||||
 Authors Authors | Puskar R / Truong CD / Swain K / Li S / Cheng K-W / Wang TY / Poh Y-P / Liu H / Chou T-F / Nannenga B / Chiu P-L | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Molecular asymmetry of a photosynthetic supercomplex from green sulfur bacteria. Authors: Ryan Puskar / Chloe Du Truong / Kyle Swain / Saborni Chowdhury / Ka-Yi Chan / Shan Li / Kai-Wen Cheng / Ting Yu Wang / Yu-Ping Poh / Yuval Mazor / Haijun Liu / Tsui-Fen Chou / Brent L Nannenga / Po-Lin Chiu /  Abstract: The photochemical reaction center (RC) features a dimeric architecture for charge separation across the membrane. In green sulfur bacteria (GSB), the trimeric Fenna-Matthews-Olson (FMO) complex ...The photochemical reaction center (RC) features a dimeric architecture for charge separation across the membrane. In green sulfur bacteria (GSB), the trimeric Fenna-Matthews-Olson (FMO) complex mediates the transfer of light energy from the chlorosome antenna complex to the RC. Here we determine the structure of the photosynthetic supercomplex from the GSB Chlorobaculum tepidum using single-particle cryogenic electron microscopy (cryo-EM) and identify the cytochrome c subunit (PscC), two accessory protein subunits (PscE and PscF), a second FMO trimeric complex, and a linker pigment between FMO and the RC core. The protein subunits that are assembled with the symmetric RC core generate an asymmetric photosynthetic supercomplex. One linker bacteriochlorophyll (BChl) is located in one of the two FMO-PscA interfaces, leading to differential efficiencies of the two energy transfer branches. The two FMO trimeric complexes establish two different binding interfaces with the RC cytoplasmic surface, driven by the associated accessory subunits. This structure of the GSB photosynthetic supercomplex provides mechanistic insight into the light excitation energy transfer routes and a possible evolutionary transition intermediate of the bacterial photosynthetic supercomplex from the primitive homodimeric RC. | |||||||||
| History |
|
- Structure visualization
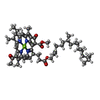
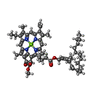
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26471.map.gz emd_26471.map.gz | 168.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26471-v30.xml emd-26471-v30.xml emd-26471.xml emd-26471.xml | 28.9 KB 28.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_26471.png emd_26471.png | 68.1 KB | ||
| Filedesc metadata |  emd-26471.cif.gz emd-26471.cif.gz | 7.9 KB | ||
| Others |  emd_26471_half_map_1.map.gz emd_26471_half_map_1.map.gz emd_26471_half_map_2.map.gz emd_26471_half_map_2.map.gz | 165.3 MB 165.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26471 http://ftp.pdbj.org/pub/emdb/structures/EMD-26471 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26471 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26471 | HTTPS FTP |
-Related structure data
| Related structure data |  7uebMC  7ueaC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_26471.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26471.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | RC-FMO2 map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.04 Å | ||||||||||||||||||||||||||||||||||||
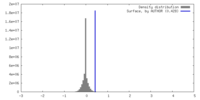
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: RC-FMO2 half map 1
| File | emd_26471_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | RC-FMO2 half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
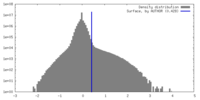
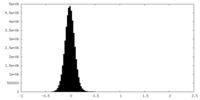
| Density Histograms |
-Half map: RC-FMO2 half map 2
| File | emd_26471_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | RC-FMO2 half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
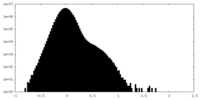
| Density Histograms |
- Sample components
Sample components
+Entire : Photosynthetic assembly (RC-FMO2)
+Supramolecule #1: Photosynthetic assembly (RC-FMO2)
+Macromolecule #1: Photosystem P840 reaction center, large subunit
+Macromolecule #2: Photosystem P840 reaction center iron-sulfur protein
+Macromolecule #3: Cytochrome c
+Macromolecule #4: P840 reaction center 17 kDa protein
+Macromolecule #5: PscE
+Macromolecule #6: PscF
+Macromolecule #7: Bacteriochlorophyll a protein
+Macromolecule #8: Bacteriochlorophyll A isomer
+Macromolecule #9: Chlorophyll A ester
+Macromolecule #10: BACTERIOCHLOROPHYLL A
+Macromolecule #11: [(2R,3S,4S,5R,6R)-6-[(10E,12E,14E)-2,6,10,14,19,23-hexamethyl-25-...
+Macromolecule #12: 2-[(1E,3E,5E,7E,9E,11E,13E,15E,17E,19E)-3,7,12,16,20,24-hexamethy...
+Macromolecule #13: 1,2-DIPALMITOYL-PHOSPHATIDYL-GLYCEROLE
+Macromolecule #14: 1,2-DISTEAROYL-MONOGALACTOSYL-DIGLYCERIDE
+Macromolecule #15: IRON/SULFUR CLUSTER
+Macromolecule #16: CALCIUM ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.1 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| ||||||||||||
| Grid | Model: C-flat-2/1 / Material: COPPER / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 15 sec. | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Instrument: HOMEMADE PLUNGER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number real images: 32898 / Average exposure time: 6.0 sec. / Average electron dose: 45.4 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Calibrated magnification: 47259 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)