[English] 日本語
 Yorodumi
Yorodumi- EMDB-25994: Structure of the rabbit 80S ribosome stalled on a 2-TMD Rhodopsin... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the rabbit 80S ribosome stalled on a 2-TMD Rhodopsin intermediate in complex with the multipass translocon | ||||||||||||
 Map data Map data | Reference map | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Ribosome / Membrane protein / translocon | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationmulti-pass transmembrane protein insertion into ER membrane / multi-pass translocon complex / Insertion of tail-anchored proteins into the endoplasmic reticulum membrane / endoplasmic reticulum Sec complex / pronephric nephron development / rough endoplasmic reticulum membrane / cotranslational protein targeting to membrane / Sec61 translocon complex / protein localization to nuclear inner membrane / protein insertion into ER membrane ...multi-pass transmembrane protein insertion into ER membrane / multi-pass translocon complex / Insertion of tail-anchored proteins into the endoplasmic reticulum membrane / endoplasmic reticulum Sec complex / pronephric nephron development / rough endoplasmic reticulum membrane / cotranslational protein targeting to membrane / Sec61 translocon complex / protein localization to nuclear inner membrane / protein insertion into ER membrane / protein-transporting ATPase activity / protein targeting to ER / SRP-dependent cotranslational protein targeting to membrane, translocation / signal sequence binding / post-translational protein targeting to membrane, translocation / endoplasmic reticulum calcium ion homeostasis / ER overload response / transmembrane protein transporter activity / protein folding chaperone complex / regulation of signal transduction / ubiquitin ligase inhibitor activity / positive regulation of signal transduction by p53 class mediator / protein-RNA complex assembly / ERAD pathway / rough endoplasmic reticulum / protein folding chaperone / MDM2/MDM4 family protein binding / guanyl-nucleotide exchange factor activity / ribosomal large subunit biogenesis / maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / post-embryonic development / antimicrobial humoral immune response mediated by antimicrobial peptide / heparin binding / large ribosomal subunit / ribosome binding / 5S rRNA binding / ribosomal large subunit assembly / large ribosomal subunit rRNA binding / nuclear membrane / killing of cells of another organism / cytosolic large ribosomal subunit / defense response to Gram-negative bacterium / cytoplasmic translation / tRNA binding / postsynaptic density / rRNA binding / structural constituent of ribosome / ribosome / translation / ribonucleoprotein complex / mRNA binding / calcium ion binding / synapse / endoplasmic reticulum membrane / nucleolus / endoplasmic reticulum / RNA binding / zinc ion binding / membrane / nucleus / plasma membrane / cytoplasm / cytosol Similarity search - Function | ||||||||||||
| Biological species |   | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.25 Å | ||||||||||||
 Authors Authors | Kim MK / Lewis AJO / Keenan RJ / Hegde RS | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nature / Year: 2022 Journal: Nature / Year: 2022Title: Mechanism of an intramembrane chaperone for multipass membrane proteins. Authors: Luka Smalinskaitė / Min Kyung Kim / Aaron J O Lewis / Robert J Keenan / Ramanujan S Hegde /   Abstract: Multipass membrane proteins play numerous roles in biology and include receptors, transporters, ion channels and enzymes. How multipass proteins are co-translationally inserted and folded at the ...Multipass membrane proteins play numerous roles in biology and include receptors, transporters, ion channels and enzymes. How multipass proteins are co-translationally inserted and folded at the endoplasmic reticulum is not well understood. The prevailing model posits that each transmembrane domain (TMD) of a multipass protein successively passes into the lipid bilayer through a front-side lateral gate of the Sec61 protein translocation channel. The PAT complex, an intramembrane chaperone comprising Asterix and CCDC47, engages early TMDs of multipass proteins to promote their biogenesis by an unknown mechanism. Here, biochemical and structural analysis of intermediates during multipass protein biogenesis showed that the nascent chain is not engaged with Sec61, which is occluded and latched closed by CCDC47. Instead, Asterix binds to and redirects the substrate to a location behind Sec61, where the PAT complex contributes to a multipass translocon surrounding a semi-enclosed, lipid-filled cavity. Detection of multiple TMDs in this cavity after their emergence from the ribosome suggests that multipass proteins insert and fold behind Sec61. Accordingly, biogenesis of several multipass proteins was unimpeded by inhibitors of the Sec61 lateral gate. These findings elucidate the mechanism of an intramembrane chaperone and suggest a new framework for multipass membrane protein biogenesis at the endoplasmic reticulum. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25994.map.gz emd_25994.map.gz | 211.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25994-v30.xml emd-25994-v30.xml emd-25994.xml emd-25994.xml | 71.4 KB 71.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_25994.png emd_25994.png | 94.9 KB | ||
| Filedesc metadata |  emd-25994.cif.gz emd-25994.cif.gz | 15.8 KB | ||
| Others |  emd_25994_additional_1.map.gz emd_25994_additional_1.map.gz emd_25994_additional_2.map.gz emd_25994_additional_2.map.gz | 213.1 MB 211.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25994 http://ftp.pdbj.org/pub/emdb/structures/EMD-25994 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25994 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25994 | HTTPS FTP |
-Related structure data
| Related structure data |  7tm3MC  7tutC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_25994.map.gz / Format: CCP4 / Size: 266.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25994.map.gz / Format: CCP4 / Size: 266.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reference map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.33981 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Focused map (PAT complex)
| File | emd_25994_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Focused map (PAT complex) | ||||||||||||
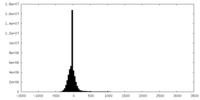
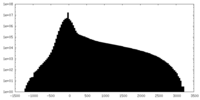
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Focused map (TMEM147 complex)
| File | emd_25994_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Focused map (TMEM147 complex) | ||||||||||||
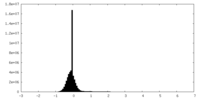
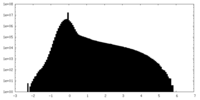
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : 80S ribosome translating a stalled, two-TMD nascent chain (derive...
+Supramolecule #1: 80S ribosome translating a stalled, two-TMD nascent chain (derive...
+Macromolecule #1: uL2
+Macromolecule #2: Nascent chain
+Macromolecule #3: uL4
+Macromolecule #4: uL18
+Macromolecule #5: eL6
+Macromolecule #6: uL30
+Macromolecule #7: eL8
+Macromolecule #8: uL6
+Macromolecule #9: uL16
+Macromolecule #10: uL5
+Macromolecule #11: eL13
+Macromolecule #12: eL14
+Macromolecule #13: eL15
+Macromolecule #14: uL13
+Macromolecule #15: uL22
+Macromolecule #16: eL18
+Macromolecule #17: eL19
+Macromolecule #18: eL20
+Macromolecule #19: eL21
+Macromolecule #20: eL22
+Macromolecule #21: uL14
+Macromolecule #22: eL24
+Macromolecule #23: uL23
+Macromolecule #24: uL24
+Macromolecule #25: eL27
+Macromolecule #26: uL15
+Macromolecule #27: eL29
+Macromolecule #28: eL30
+Macromolecule #29: eL31
+Macromolecule #30: eL32
+Macromolecule #31: eL33
+Macromolecule #32: eL34
+Macromolecule #33: uL29
+Macromolecule #34: eL36
+Macromolecule #35: eL37
+Macromolecule #36: eL38
+Macromolecule #37: eL39
+Macromolecule #38: eL40
+Macromolecule #39: eL41
+Macromolecule #40: eL42
+Macromolecule #41: eL43
+Macromolecule #43: eL28
+Macromolecule #46: uL3
+Macromolecule #47: Protein transport protein Sec61 subunit alpha isoform 1
+Macromolecule #48: Protein transport protein Sec61 subunit beta
+Macromolecule #49: Protein transport protein Sec61 gamma
+Macromolecule #50: Coiled-coil domain containing 47
+Macromolecule #51: PAT complex subunit Asterix
+Macromolecule #52: Nicalin
+Macromolecule #53: Transmembrane protein 147
+Macromolecule #42: P-site tRNA
+Macromolecule #44: 5S ribosomal RNA
+Macromolecule #45: 5.8S ribosomal RNA
+Macromolecule #54: 28S ribosomal RNA
+Macromolecule #55: MAGNESIUM ION
+Macromolecule #56: ZINC ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Details: 50 mM HEPES-KOH, pH 7.5, 200 mM KOAc, 5 mM Mg(OAc)2, 0.25% digitonin, 50 mM biotin |
|---|---|
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GRAPHENE OXIDE / Mesh: 300 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV Details: blot for 4 sec with Whatman filter papers at a blot force of -15 before plunging. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 1 / Number real images: 17540 / Average electron dose: 54.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.7 µm / Nominal defocus min: 1.9000000000000001 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: OTHER |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.25 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 1665551 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-7tm3: |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)