[English] 日本語
 Yorodumi
Yorodumi- EMDB-25792: Cryo-EM structure of the spike of SARS-CoV-2 Omicron variant of c... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-25792 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the spike of SARS-CoV-2 Omicron variant of concern | |||||||||
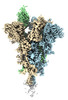
 Map data Map data | cryo-EM map for the SARS-CoV-2 Omicron spike protein | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | SARS-CoV-2 / spike / Omicron / variant of concern / VIRAL PROTEIN | |||||||||
| Function / homology | Surface glycoprotein Function and homology information Function and homology information | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.29 Å | |||||||||
 Authors Authors | Zhou T / Tsybovsky T | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: bioRxiv / Year: 2021 Journal: bioRxiv / Year: 2021Title: Antibodies with potent and broad neutralizing activity against antigenically diverse and highly transmissible SARS-CoV-2 variants. Authors: Lingshu Wang / Tongqing Zhou / Yi Zhang / Eun Sung Yang / Chaim A Schramm / Wei Shi / Amarendra Pegu / Olamide K Oloninyi / Amy Ransier / Samuel Darko / Sandeep R Narpala / Christian Hatcher ...Authors: Lingshu Wang / Tongqing Zhou / Yi Zhang / Eun Sung Yang / Chaim A Schramm / Wei Shi / Amarendra Pegu / Olamide K Oloninyi / Amy Ransier / Samuel Darko / Sandeep R Narpala / Christian Hatcher / David R Martinez / Yaroslav Tsybovsky / Emily Phung / Olubukola M Abiona / Evan M Cale / Lauren A Chang / Kizzmekia S Corbett / Anthony T DiPiazza / Ingelise J Gordon / Kwanyee Leung / Tracy Liu / Rosemarie D Mason / Alexandra Nazzari / Laura Novik / Adam S Olia / Nicole A Doria-Rose / Tyler Stephens / Christopher D Stringham / Chloe Adrienna Talana / I-Ting Teng / Danielle Wagner / Alicia T Widge / Baoshan Zhang / Mario Roederer / Julie E Ledgerwood / Tracy J Ruckwardt / Martin R Gaudinski / Ralph S Baric / Barney S Graham / Adrian B McDermott / Daniel C Douek / Peter D Kwong / John R Mascola / Nancy J Sullivan / John Misasi /  Abstract: The emergence of highly transmissible SARS-CoV-2 variants of concern (VOC) that are resistant to therapeutic antibodies highlights the need for continuing discovery of broadly reactive antibodies. We ...The emergence of highly transmissible SARS-CoV-2 variants of concern (VOC) that are resistant to therapeutic antibodies highlights the need for continuing discovery of broadly reactive antibodies. We identify four receptor-binding domain targeting antibodies from three early-outbreak convalescent donors with potent neutralizing activity against 12 variants including the B.1.1.7 and B.1.351 VOCs. Two of them are ultrapotent, with sub-nanomolar neutralization titers (IC50 <0.0006 to 0.0102 μ g/mL; IC80 < 0.0006 to 0.0251 μ g/mL). We define the structural and functional determinants of binding for all four VOC-targeting antibodies, and show that combinations of two antibodies decrease the in vitro generation of escape mutants, suggesting potential means to mitigate resistance development. These results define the basis of therapeutic cocktails against VOCs and suggest that targeted boosting of existing immunity may increase vaccine breadth against VOCs. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25792.map.gz emd_25792.map.gz | 121.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25792-v30.xml emd-25792-v30.xml emd-25792.xml emd-25792.xml | 21.2 KB 21.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_25792.png emd_25792.png | 104.8 KB | ||
| Masks |  emd_25792_msk_1.map emd_25792_msk_1.map | 244.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-25792.cif.gz emd-25792.cif.gz | 7 KB | ||
| Others |  emd_25792_additional_1.map.gz emd_25792_additional_1.map.gz emd_25792_half_map_1.map.gz emd_25792_half_map_1.map.gz emd_25792_half_map_2.map.gz emd_25792_half_map_2.map.gz | 230 MB 226.4 MB 226.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25792 http://ftp.pdbj.org/pub/emdb/structures/EMD-25792 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25792 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25792 | HTTPS FTP |
-Related structure data
| Related structure data |  7tb4MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_25792.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25792.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cryo-EM map for the SARS-CoV-2 Omicron spike protein | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.855 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_25792_msk_1.map emd_25792_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Sharpened cryo-EM map for the SARS-CoV-2 Omicron spike protein
| File | emd_25792_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened cryo-EM map for the SARS-CoV-2 Omicron spike protein | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: cryo-EM half map for the SARS-CoV-2 Omicron spike protein
| File | emd_25792_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cryo-EM half map for the SARS-CoV-2 Omicron spike protein | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: cryo-EM half map for the SARS-CoV-2 Omicron spike protein
| File | emd_25792_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cryo-EM half map for the SARS-CoV-2 Omicron spike protein | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : SARS-CoV-2 spike of the Omicron Variant of Concern
| Entire | Name: SARS-CoV-2 spike of the Omicron Variant of Concern |
|---|---|
| Components |
|
-Supramolecule #1: SARS-CoV-2 spike of the Omicron Variant of Concern
| Supramolecule | Name: SARS-CoV-2 spike of the Omicron Variant of Concern / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 414 KDa |
-Macromolecule #1: Surface glycoprotein
| Macromolecule | Name: Surface glycoprotein / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 132.465094 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QCVNLTTRTQ LPPAYTNSFT RGVYYPDKVF RSSVLHSTQD LFLPFFSNVT WFHVISGTNG TKRFDNPVLP FNDGVYFASI EKSNIIRGW IFGTTLDSKT QSLLIVNNAT NVVIKVCEFQ FCNDPFLDHK NNKSWMESEF RVYSSANNCT FEYVSQPFLM D LEGKQGNF ...String: QCVNLTTRTQ LPPAYTNSFT RGVYYPDKVF RSSVLHSTQD LFLPFFSNVT WFHVISGTNG TKRFDNPVLP FNDGVYFASI EKSNIIRGW IFGTTLDSKT QSLLIVNNAT NVVIKVCEFQ FCNDPFLDHK NNKSWMESEF RVYSSANNCT FEYVSQPFLM D LEGKQGNF KNLREFVFKN IDGYFKIYSK HTPIIVREPE DLPQGFSALE PLVDLPIGIN ITRFQTLLAL HRSYLTPGDS SS GWTAGAA AYYVGYLQPR TFLLKYNENG TITDAVDCAL DPLSETKCTL KSFTVEKGIY QTSNFRVQPT ESIVRFPNIT NLC PFDEVF NATRFASVYA WNRKRISNCV ADYSVLYNLA PFFTFKCYGV SPTKLNDLCF TNVYADSFVI RGDEVRQIAP GQTG NIADY NYKLPDDFTG CVIAWNSNKL DSKVSGNYNY LYRLFRKSNL KPFERDISTE IYQAGNKPCN GVAGFNCYFP LRSYS FRPT YGVGHQPYRV VVLSFELLHA PATVCGPKKS TNLVKNKCVN FNFNGLKGTG VLTESNKKFL PFQQFGRDIA DTTDAV RDP QTLEILDITP CSFGGVSVIT PGTNTSNQVA VLYQGVNCTE VPVAIHADQL TPTWRVYSTG SNVFQTRAGC LIGAEYV NN SYECDIPIGA GICASYQTQT KSHGSASSVA SQSIIAYTMS LGAENSVAYS NNSIAIPTNF TISVTTEILP VSMTKTSV D CTMYICGDST ECSNLLLQYG SFCTQLKRAL TGIAVEQDKN TQEVFAQVKQ IYKTPPIKYF GGFNFSQILP DPSKPSKRS FIEDLLFNKV TLADAGFIKQ YGDCLGDIAA RDLICAQKFK GLTVLPPLLT DEMIAQYTSA LLAGTITSGW TFGAGAALQI PFAMQMAYR FNGIGVTQNV LYENQKLIAN QFNSAIGKIQ DSLSSTASAL GKLQDVVNHN AQALNTLVKQ LSSKFGAISS V LNDIFSRL DPPEAEVQID RLITGRLQSL QTYVTQQLIR AAEIRASANL AATKMSECVL GQSKRVDFCG KGYHLMSFPQ SA PHGVVFL HVTYVPAQEK NFTTAPAICH DGKAHFPREG VFVSNGTHWF VTQRNFYEPQ IITTDNTFVS GNCDVVIGIV NNT VYDPLQ PELDSFKEEL DKYFKNHTSP DVDLGDISGI NASVVNIQKE IDRLNEVAKN LNESLIDLQE LGKYEQ UniProtKB: Surface glycoprotein |
-Macromolecule #3: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 3 / Number of copies: 29 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Details: 100 mM HEPES, 150 mM NaCl |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: Blot for 2-3.5 seconds before plugging.. |
| Details | Complex at 0.5 mg/mL concentration in the buffer |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: OTHER / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: Details: SARS-CoV-2 receptor-binding domain in complex with antibody A23-58.1 |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.29 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: cryoSPARC (ver. 2.15) / Number images used: 266434 |
| Initial angle assignment | Type: NOT APPLICABLE |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)