[English] 日本語
 Yorodumi
Yorodumi- EMDB-25210: Structure of human SARS-CoV-2 spike glycoprotein trimer bound by ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-25210 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of human SARS-CoV-2 spike glycoprotein trimer bound by neutralizing antibody C1C-A3 Fab (variable region) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | COVID-19 / SARS-CoV-2 / neutralizing antibody / neutralization escape / IMMUNE SYSTEM / VIRAL PROTEIN-IMMUNE SYSTEM complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular region / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / positive regulation of viral entry into host cell ...symbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular region / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / positive regulation of viral entry into host cell / membrane fusion / Attachment and Entry / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / entry receptor-mediated virion attachment to host cell / receptor-mediated virion attachment to host cell / host cell surface receptor binding / symbiont-mediated suppression of host innate immune response / endocytosis involved in viral entry into host cell / receptor ligand activity / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / symbiont entry into host cell / virion attachment to host cell / host cell plasma membrane / SARS-CoV-2 activates/modulates innate and adaptive immune responses / virion membrane / membrane / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||
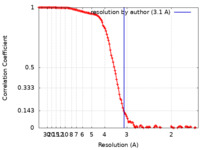
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Pan J / Abraham J | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2022 Journal: Science / Year: 2022Title: Structural basis for continued antibody evasion by the SARS-CoV-2 receptor binding domain. Authors: Katherine G Nabel / Sarah A Clark / Sundaresh Shankar / Junhua Pan / Lars E Clark / Pan Yang / Adrian Coscia / Lindsay G A McKay / Haley H Varnum / Vesna Brusic / Nicole V Tolan / Guohai ...Authors: Katherine G Nabel / Sarah A Clark / Sundaresh Shankar / Junhua Pan / Lars E Clark / Pan Yang / Adrian Coscia / Lindsay G A McKay / Haley H Varnum / Vesna Brusic / Nicole V Tolan / Guohai Zhou / Michaël Desjardins / Sarah E Turbett / Sanjat Kanjilal / Amy C Sherman / Anand Dighe / Regina C LaRocque / Edward T Ryan / Casey Tylek / Joel F Cohen-Solal / Anhdao T Darcy / Davide Tavella / Anca Clabbers / Yao Fan / Anthony Griffiths / Ivan R Correia / Jane Seagal / Lindsey R Baden / Richelle C Charles / Jonathan Abraham /   Abstract: Many studies have examined the impact of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) variants on neutralizing antibody activity after they have become dominant strains. Here, we ...Many studies have examined the impact of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) variants on neutralizing antibody activity after they have become dominant strains. Here, we evaluate the consequences of further viral evolution. We demonstrate mechanisms through which the SARS-CoV-2 receptor binding domain (RBD) can tolerate large numbers of simultaneous antibody escape mutations and show that pseudotypes containing up to seven mutations, as opposed to the one to three found in previously studied variants of concern, are more resistant to neutralization by therapeutic antibodies and serum from vaccine recipients. We identify an antibody that binds the RBD core to neutralize pseudotypes for all tested variants but show that the RBD can acquire an N-linked glycan to escape neutralization. Our findings portend continued emergence of escape variants as SARS-CoV-2 adapts to humans. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25210.map.gz emd_25210.map.gz | 167.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25210-v30.xml emd-25210-v30.xml emd-25210.xml emd-25210.xml | 23.2 KB 23.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_25210_fsc.xml emd_25210_fsc.xml | 12.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_25210.png emd_25210.png | 144 KB | ||
| Filedesc metadata |  emd-25210.cif.gz emd-25210.cif.gz | 8.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25210 http://ftp.pdbj.org/pub/emdb/structures/EMD-25210 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25210 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25210 | HTTPS FTP |
-Related structure data
| Related structure data |  7sn3MC  7sn0C  7sn1C  7sn2C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_25210.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25210.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.825 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : SARS-CoV-2 spike protein trimer in complex with human patient ant...
| Entire | Name: SARS-CoV-2 spike protein trimer in complex with human patient antibody C1C-A3 Fab |
|---|---|
| Components |
|
-Supramolecule #1: SARS-CoV-2 spike protein trimer in complex with human patient ant...
| Supramolecule | Name: SARS-CoV-2 spike protein trimer in complex with human patient antibody C1C-A3 Fab type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 Details: Concentration of the SARS-CoV-2 spike protein (trimeric) and human patient antibody C1C-A3 Fab is 0.6 and 0.3 mg/mL in the final complex sample in buffer consisting of 150 mM NaCl and 25 mM Tris-HCl, pH 7.5 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 50 KDa |
-Supramolecule #2: SARS-CoV-2 spike protein
| Supramolecule | Name: SARS-CoV-2 spike protein / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 Details: the SARS-Cov-2 spike protein is a trimeric protein of a calculated amino acid molecular weight of 418176 Da, with 23 N-linked glycans. Total molecular weight is 556176 Da assuming each glycan is 2000 Da. |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: C1C-A3 Fab (variable region)
| Supramolecule | Name: C1C-A3 Fab (variable region) / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2-#3 Details: Fab of C1C-A3 antibody recombinantly expressed using sequence sequenced from single B cells from human patient peripheral blood sample |
|---|
-Macromolecule #1: Spike glycoprotein
| Macromolecule | Name: Spike glycoprotein / type: protein_or_peptide / ID: 1 Details: amino acids 1-15 are the endogenous signal peptide, 16-1208 are SARS-CoV-2 spike protein residues 16-1208, 1209-1210, 1237-1239, and 1255-1258 are linkers, 1211-1236 are T4 fibritinn foldon, ...Details: amino acids 1-15 are the endogenous signal peptide, 16-1208 are SARS-CoV-2 spike protein residues 16-1208, 1209-1210, 1237-1239, and 1255-1258 are linkers, 1211-1236 are T4 fibritinn foldon, 1240-1254 are the avi-tag for biotinylation, 1259-1265 are the TEV protease cleavage site, 1266-1273 are the 8xHis tag; mutations include R682G, R683S, R685S, F817P, A892P, A899P, A942P, K986P, V987P Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 141.192203 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MFVFLVLLPL VSSQCVNLTT RTQLPPAYTN SFTRGVYYPD KVFRSSVLHS TQDLFLPFFS NVTWFHAIHV SGTNGTKRFD NPVLPFNDG VYFASTEKSN IIRGWIFGTT LDSKTQSLLI VNNATNVVIK VCEFQFCNDP FLGVYYHKNN KSWMESEFRV Y SSANNCTF ...String: MFVFLVLLPL VSSQCVNLTT RTQLPPAYTN SFTRGVYYPD KVFRSSVLHS TQDLFLPFFS NVTWFHAIHV SGTNGTKRFD NPVLPFNDG VYFASTEKSN IIRGWIFGTT LDSKTQSLLI VNNATNVVIK VCEFQFCNDP FLGVYYHKNN KSWMESEFRV Y SSANNCTF EYVSQPFLMD LEGKQGNFKN LREFVFKNID GYFKIYSKHT PINLVRDLPQ GFSALEPLVD LPIGINITRF QT LLALHRS YLTPGDSSSG WTAGAAAYYV GYLQPRTFLL KYNENGTITD AVDCALDPLS ETKCTLKSFT VEKGIYQTSN FRV QPTESI VRFPNITNLC PFGEVFNATR FASVYAWNRK RISNCVADYS VLYNSASFST FKCYGVSPTK LNDLCFTNVY ADSF VIRGD EVRQIAPGQT GKIADYNYKL PDDFTGCVIA WNSNNLDSKV GGNYNYLYRL FRKSNLKPFE RDISTEIYQA GSTPC NGVE GFNCYFPLQS YGFQPTNGVG YQPYRVVVLS FELLHAPATV CGPKKSTNLV KNKCVNFNFN GLTGTGVLTE SNKKFL PFQ QFGRDIADTT DAVRDPQTLE ILDITPCSFG GVSVITPGTN TSNQVAVLYQ DVNCTEVPVA IHADQLTPTW RVYSTGS NV FQTRAGCLIG AEHVNNSYEC DIPIGAGICA SYQTQTNSPG SASSVASQSI IAYTMSLGAE NSVAYSNNSI AIPTNFTI S VTTEILPVSM TKTSVDCTMY ICGDSTECSN LLLQYGSFCT QLNRALTGIA VEQDKNTQEV FAQVKQIYKT PPIKDFGGF NFSQILPDPS KPSKRSPIED LLFNKVTLAD AGFIKQYGDC LGDIAARDLI CAQKFNGLTV LPPLLTDEMI AQYTSALLAG TITSGWTFG AGPALQIPFP MQMAYRFNGI GVTQNVLYEN QKLIANQFNS AIGKIQDSLS STPSALGKLQ DVVNQNAQAL N TLVKQLSS NFGAISSVLN DILSRLDPPE AEVQIDRLIT GRLQSLQTYV TQQLIRAAEI RASANLAATK MSECVLGQSK RV DFCGKGY HLMSFPQSAP HGVVFLHVTY VPAQEKNFTT APAICHDGKA HFPREGVFVS NGTHWFVTQR NFYEPQIITT DNT FVSGNC DVVIGIVNNT VYDPLQPELD SFKEELDKYF KNHTSPDVDL GDISGINASV VNIQKEIDRL NEVAKNLNES LIDL QELGK YEQSGYIPEA PRDGQAYVRK DGEWVLLSTF LSAGGLNDIF EAQKIEWHEG ASGENLYFQG HHHHHHHH UniProtKB: Spike glycoprotein |
-Macromolecule #2: neutralizing antibody C1C-A3 heavy chain variable region
| Macromolecule | Name: neutralizing antibody C1C-A3 heavy chain variable region type: protein_or_peptide / ID: 2 Details: amino acids 1-20 are the human tPA signal peptide, 21-22 are a linker, 23-250 are the C1C-A3 Fab heavy chain residues 1-213 (note insertions in CDR loops; numbering should be according to the PDB file). Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 27.022607 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MKRGLCCVLL LCGAVFVSPS ASQVQLVESG GGVVQPGRSL RLSCAASGFT FSSYGMHWVR QAPGKGLEWV AVIWYDGTNT YYADSVKGR FTISRDNSKN TLYLQMNSLR AEDTAVYYCA RDLAYRDYVW RYFDLWGRGT LVTVSGASTK GPSVFPLAPS S KSTSGGTA ...String: MKRGLCCVLL LCGAVFVSPS ASQVQLVESG GGVVQPGRSL RLSCAASGFT FSSYGMHWVR QAPGKGLEWV AVIWYDGTNT YYADSVKGR FTISRDNSKN TLYLQMNSLR AEDTAVYYCA RDLAYRDYVW RYFDLWGRGT LVTVSGASTK GPSVFPLAPS S KSTSGGTA ALGCLVKDYF PEPVTVSWNS GALTSGVHTF PAVLQSSGLY SLSSVVTVPS SSLGTQTYIC NVNHKPSNTK VD KRVEPKS CDR |
-Macromolecule #3: neutralizing antibody C1C-A3 light chain variable region
| Macromolecule | Name: neutralizing antibody C1C-A3 light chain variable region type: protein_or_peptide / ID: 3 Details: amino acids 1-20 are the human tPA signal peptide, 21-22 are a linker, 23-238 are the C1C-A3 Fab light chain residues 1-212 (note insertions in CDR loops; numbering should be according to the PDB file). Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 25.896107 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MKRGLCCVLL LCGAVFVSPS ASEIVLTQSP ATLSLSPGER ATLSCRASQS VSTYLAWYQQ KFGQAPRLLI YDASNRATGI PARFSGSGS GTDFTLTISS LEPEDFAVYY CQCRSNWPPG ITFGQGTRLE IKRTVAAPSV FIFPPSDEQL KSGTASVVCL L NNFYPREA ...String: MKRGLCCVLL LCGAVFVSPS ASEIVLTQSP ATLSLSPGER ATLSCRASQS VSTYLAWYQQ KFGQAPRLLI YDASNRATGI PARFSGSGS GTDFTLTISS LEPEDFAVYY CQCRSNWPPG ITFGQGTRLE IKRTVAAPSV FIFPPSDEQL KSGTASVVCL L NNFYPREA KVQWKVDNAL QSGNSQESVT EQDSKDSTYS LSSTLTLSKA DYEKHKVYAC EVTHQGLSSP VTKSFNRGEC |
-Macromolecule #5: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 5 / Number of copies: 45 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.9 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| |||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 2 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 20 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.039 kPa / Details: glow discharge done at 20 mA in Pelco easiGlow | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 293 K / Instrument: FEI VITROBOT MARK IV / Details: blotting time 4-6 seconds. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 77.0 K / Max: 77.0 K |
| Details | Cs-corrected, energy filtered |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Number grids imaged: 1 / Number real images: 4583 / Average exposure time: 0.038 sec. / Average electron dose: 56.6 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated magnification: 60606 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: OTHER / Overall B value: 77.28 / Target criteria: correlation coefficient |
|---|---|
| Output model |  PDB-7sn3: |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)