[English] 日本語
 Yorodumi
Yorodumi- EMDB-19433: Cryo-em structure of the rat Multidrug resistance-associated prot... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-em structure of the rat Multidrug resistance-associated protein 2 (rMrp2) in complex with probenecid | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Multidrug resistance-associated protein 2 (rMrp2) in an autoinhibited state (nucleotide-free) / TRANSPORT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationmercury ion transport / benzylpenicillin metabolic process / Aspirin ADME / Paracetamol ADME / Atorvastatin ADME / canalicular bile acid transport / intracellular canaliculus / antibiotic metabolic process / bilirubin transmembrane transporter activity / bilirubin transport ...mercury ion transport / benzylpenicillin metabolic process / Aspirin ADME / Paracetamol ADME / Atorvastatin ADME / canalicular bile acid transport / intracellular canaliculus / antibiotic metabolic process / bilirubin transmembrane transporter activity / bilirubin transport / xenobiotic export from cell / Heme degradation / response to antineoplastic agent / leukotriene transport / detoxification of mercury ion / thyroid hormone transport / ABC-family proteins mediated transport / regulation of bile acid secretion / prostaglandin transport / ABC-type glutathione-S-conjugate transporter / ABC-type glutathione S-conjugate transporter activity / intracellular chloride ion homeostasis / organic anion transport / organic anion transmembrane transporter activity / xenobiotic transmembrane transport / xenobiotic transport across blood-brain barrier / transepithelial transport / xenobiotic detoxification by transmembrane export across the plasma membrane / intercellular canaliculus / ABC-type xenobiotic transporter / response to arsenic-containing substance / bile acid and bile salt transport / cellular response to interleukin-6 / ABC-type xenobiotic transporter activity / Translocases; Catalysing the translocation of other compounds; Linked to the hydrolysis of a nucleoside triphosphate / response to glucagon / response to steroid hormone / bile acid signaling pathway / xenobiotic transmembrane transporter activity / cellular response to interleukin-1 / ATPase-coupled transmembrane transporter activity / xenobiotic catabolic process / cellular response to dexamethasone stimulus / female pregnancy / brush border membrane / response to organic cyclic compound / transmembrane transport / response to estrogen / cellular response to xenobiotic stimulus / response to estradiol / cellular response to tumor necrosis factor / cellular response to lipopolysaccharide / response to oxidative stress / response to lipopolysaccharide / apical plasma membrane / response to xenobiotic stimulus / protein domain specific binding / negative regulation of gene expression / cell surface / ATP hydrolysis activity / ATP binding / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.45 Å | |||||||||
 Authors Authors | Mazza T / Beis K | |||||||||
| Funding support |  Canada, 1 items Canada, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Structural basis for the modulation of MRP2 activity by phosphorylation and drugs. Authors: Tiziano Mazza / Theodoros I Roumeliotis / Elena Garitta / David Drew / S Tamir Rashid / Cesare Indiveri / Jyoti S Choudhary / Kenneth J Linton / Konstantinos Beis /    Abstract: Multidrug resistance-associated protein 2 (MRP2/ABCC2) is a polyspecific efflux transporter of organic anions expressed in hepatocyte canalicular membranes. MRP2 dysfunction, in Dubin-Johnson ...Multidrug resistance-associated protein 2 (MRP2/ABCC2) is a polyspecific efflux transporter of organic anions expressed in hepatocyte canalicular membranes. MRP2 dysfunction, in Dubin-Johnson syndrome or by off-target inhibition, for example by the uricosuric drug probenecid, elevates circulating bilirubin glucuronide and is a cause of jaundice. Here, we determine the cryo-EM structure of rat Mrp2 (rMrp2) in an autoinhibited state and in complex with probenecid. The autoinhibited state exhibits an unusual conformation for this class of transporter in which the regulatory domain is folded within the transmembrane domain cavity. In vitro phosphorylation, mass spectrometry and transport assays show that phosphorylation of the regulatory domain relieves this autoinhibition and enhances rMrp2 transport activity. The in vitro data is confirmed in human hepatocyte-like cells, in which inhibition of endogenous kinases also reduces human MRP2 transport activity. The drug-bound state reveals two probenecid binding sites that suggest a dynamic interplay with autoinhibition. Mapping of the Dubin-Johnson mutations onto the rodent structure indicates that many may interfere with the transition between conformational states. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_19433.map.gz emd_19433.map.gz | 306.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-19433-v30.xml emd-19433-v30.xml emd-19433.xml emd-19433.xml | 16 KB 16 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_19433.png emd_19433.png | 41.1 KB | ||
| Filedesc metadata |  emd-19433.cif.gz emd-19433.cif.gz | 6.6 KB | ||
| Others |  emd_19433_half_map_1.map.gz emd_19433_half_map_1.map.gz emd_19433_half_map_2.map.gz emd_19433_half_map_2.map.gz | 301.1 MB 301.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-19433 http://ftp.pdbj.org/pub/emdb/structures/EMD-19433 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19433 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19433 | HTTPS FTP |
-Validation report
| Summary document |  emd_19433_validation.pdf.gz emd_19433_validation.pdf.gz | 1.2 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_19433_full_validation.pdf.gz emd_19433_full_validation.pdf.gz | 1.2 MB | Display | |
| Data in XML |  emd_19433_validation.xml.gz emd_19433_validation.xml.gz | 16.8 KB | Display | |
| Data in CIF |  emd_19433_validation.cif.gz emd_19433_validation.cif.gz | 19.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19433 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19433 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19433 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19433 | HTTPS FTP |
-Related structure data
| Related structure data |  8rq4MC  8rq3C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_19433.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_19433.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.65 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_19433_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_19433_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cryo-em structure of the rat Multidrug resistance-associated prot...
| Entire | Name: Cryo-em structure of the rat Multidrug resistance-associated protein 2 (rMrp2) in complex with probenecid |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-em structure of the rat Multidrug resistance-associated prot...
| Supramolecule | Name: Cryo-em structure of the rat Multidrug resistance-associated protein 2 (rMrp2) in complex with probenecid type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: ATP-binding cassette sub-family C member 2
| Macromolecule | Name: ATP-binding cassette sub-family C member 2 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO EC number: Translocases; Catalysing the translocation of other compounds; Linked to the hydrolysis of a nucleoside triphosphate |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 173.571578 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MDKFCNSTFW DLSLLESPEA DLPLCFEQTV LVWIPLGFLW LLAPWQLYSV YRSRTKRSSI TKFYLAKQVF VVFLLILAAI DLSLALTED TGQATVPPVR YTNPILYLCT WLLVLAVQHS RQWCVRKNSW FLSLFWILSV LCGVFQFQTL IRALLKDSKS N MAYSYLFF ...String: MDKFCNSTFW DLSLLESPEA DLPLCFEQTV LVWIPLGFLW LLAPWQLYSV YRSRTKRSSI TKFYLAKQVF VVFLLILAAI DLSLALTED TGQATVPPVR YTNPILYLCT WLLVLAVQHS RQWCVRKNSW FLSLFWILSV LCGVFQFQTL IRALLKDSKS N MAYSYLFF VSYGFQIVLL ILTAFSGPSD STQTPSVTAS FLSSITFSWY DRTVLKGYKH PLTLEDVWDI DEGFKTRSVT SK FEAAMTK DLQKARQAFQ RRLQKSQRKP EATLHGLNKK QSQSQDVLVL EEAKKKSEKT TKDYPKSWLI KSLFKTFHVV ILK SFILKL IHDLLVFLNP QLLKLLIGFV KSSNSYVWFG YICAILMFAV TLIQSFCLQS YFQHCFVLGM CVRTTVMSSI YKKA LTLSN LARKQYTIGE TVNLMSVDSQ KLMDATNYMQ LVWSSVIQIT LSIFFLWREL GPSILAGVGV MVLLIPVNGV LATKI RNIQ VQNMKNKDKR LKIMNEILSG IKILKYFAWE PSFQEQVQGI RKKELKNLLR FGQLQSLLIF ILQITPILVS VVTFSV YVL VDSANVLNAE KAFTSITLFN ILRFPLSMLP MVTSSILQAS VSVDRLERYL GGDDLDTSAI RRVSNFDKAV KFSEASF TW DPDLEATIQD VNLDIKPGQL VAVVGTVGSG KSSLVSAMLG EMENVHGHIT IQGSTAYVPQ QSWIQNGTIK DNILFGSE Y NEKKYQQVLK ACALLPDLEI LPGGDMAEIG EKGINLSGGQ KQRVSLARAA YQDADIYILD DPLSAVDAHV GKHIFNKVV GPNGLLAGKT RIFVTHGIHF LPQVDEIVVL GKGTILEKGS YRDLLDKKGV FARNWKTFMK HSGPEGEATV NNDSEAEDDD DGLIPTMEE IPEDAASLAM RRENSLRRTL SRSSRSSSRR GKSLKNSLKI KNVNVLKEKE KEVEGQKLIK KEFVETGKVK F SIYLKYLQ AVGWWSILFI ILFYGLNNVA FIGSNLWLSA WTSDSDNLNG TNNSSSHRDM RIGVFGALGL AQGICLLIST LW SIYACRN ASKALHGQLL TNILRAPMRF FDTTPTGRIV NRFSGDISTV DDLLPQTLRS WMMCFFGIAG TLVMICMATP VFA IIIIPL SILYISVQVF YVATSRQLRR LDSVTKSPIY SHFSETVTGL PIIRAFEHQQ RFLAWNEKQI DINQKCVFSW ITSN RWLAI RLELVGNLVV FCSALLLVIY RKTLTGDVVG FVLSNALNIT QTLNWLVRMT SEAETNIVAV ERISEYINVE NEAPW VTDK RPPADWPRHG EIQFNNYQVR YRPELDLVLK GITCNIKSGE KVGVVGRTGA GKSSLTNCLF RILESAGGQI IIDGID VAS IGLHDLRERL TIIPQDPILF SGSLRMNLDP FNKYSDEEVW RALELAHLRS FVSGLQLGLL SEVTEGGDNL SIGQRQL LC LGRAVLRKSK ILVLDEATAA VDLETDSLIQ TTIRKEFSQC TVITIAHRLH TIMDSDKIMV LDNGKIVEYG SPEELLSN R GSFYLMAKEA GIENVNHTEL UniProtKB: ATP-binding cassette sub-family C member 2 |
-Macromolecule #2: CHOLESTEROL HEMISUCCINATE
| Macromolecule | Name: CHOLESTEROL HEMISUCCINATE / type: ligand / ID: 2 / Number of copies: 1 / Formula: Y01 |
|---|---|
| Molecular weight | Theoretical: 486.726 Da |
| Chemical component information |  ChemComp-Y01: |
-Macromolecule #3: 4-(dipropylsulfamoyl)benzoic acid
| Macromolecule | Name: 4-(dipropylsulfamoyl)benzoic acid / type: ligand / ID: 3 / Number of copies: 2 / Formula: RTO |
|---|---|
| Molecular weight | Theoretical: 285.359 Da |
| Chemical component information | 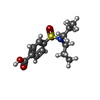 ChemComp-RTO: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 52.8 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.4 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.45 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 247763 |
| Initial angle assignment | Type: ANGULAR RECONSTITUTION |
| Final angle assignment | Type: ANGULAR RECONSTITUTION |
 Movie
Movie Controller
Controller






 Z
Z Y
Y X
X