+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | TadA/CpaF with ADP | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | pilus / secretion / ATPase / MOTOR PROTEIN | |||||||||
| Function / homology | : / Type II/IV secretion system protein / Type II/IV secretion system protein / ATP hydrolysis activity / P-loop containing nucleoside triphosphate hydrolase / Pilus assembly ATPase CpaF Function and homology information Function and homology information | |||||||||
| Biological species |  Caulobacter vibrioides NA1000 (bacteria) Caulobacter vibrioides NA1000 (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Hohl M / Low H | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Bidirectional pilus processing in the Tad pilus system motor CpaF. Authors: Michael Hohl / Emma J Banks / Max P Manley / Tung B K Le / Harry H Low /  Abstract: The bacterial tight adherence pilus system (TadPS) assembles surface pili essential for adhesion and colonisation in many human pathogens. Pilus dynamics are powered by the ATPase CpaF (TadA), which ...The bacterial tight adherence pilus system (TadPS) assembles surface pili essential for adhesion and colonisation in many human pathogens. Pilus dynamics are powered by the ATPase CpaF (TadA), which drives extension and retraction cycles in Caulobacter crescentus through an unknown mechanism. Here we use cryogenic electron microscopy and cell-based light microscopy to characterise CpaF mechanism. We show that CpaF assembles into a hexamer with C2 symmetry in different nucleotide states. Nucleotide cycling occurs through an intra-subunit clamp-like mechanism that promotes sequential conformational changes between subunits. Moreover, a comparison of the active sites with different nucleotides bound suggests a mechanism for bidirectional motion. Conserved CpaF residues, predicted to interact with platform proteins CpaG (TadB) and CpaH (TadC), are mutated in vivo to establish their role in pilus processing. Our findings provide a model for how CpaF drives TadPS pilus dynamics and have broad implications for how other ancient type 4 filament family members power pilus assembly. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_19209.map.gz emd_19209.map.gz | 161.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-19209-v30.xml emd-19209-v30.xml emd-19209.xml emd-19209.xml | 14.1 KB 14.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_19209.png emd_19209.png | 173 KB | ||
| Filedesc metadata |  emd-19209.cif.gz emd-19209.cif.gz | 5.6 KB | ||
| Others |  emd_19209_half_map_1.map.gz emd_19209_half_map_1.map.gz emd_19209_half_map_2.map.gz emd_19209_half_map_2.map.gz | 139.3 MB 138.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-19209 http://ftp.pdbj.org/pub/emdb/structures/EMD-19209 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19209 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19209 | HTTPS FTP |
-Validation report
| Summary document |  emd_19209_validation.pdf.gz emd_19209_validation.pdf.gz | 926.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_19209_full_validation.pdf.gz emd_19209_full_validation.pdf.gz | 926.2 KB | Display | |
| Data in XML |  emd_19209_validation.xml.gz emd_19209_validation.xml.gz | 15.2 KB | Display | |
| Data in CIF |  emd_19209_validation.cif.gz emd_19209_validation.cif.gz | 17.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19209 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19209 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19209 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19209 | HTTPS FTP |
-Related structure data
| Related structure data |  8rjfMC  8rkdC  8rklC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_19209.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_19209.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.829 Å | ||||||||||||||||||||||||||||||||||||
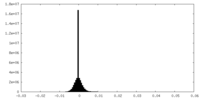
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_19209_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
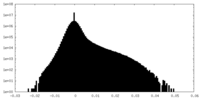
| Density Histograms |
-Half map: #1
| File | emd_19209_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
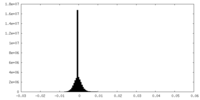
| Density Histograms |
- Sample components
Sample components
-Entire : Hexameric complex of TadA bound to ADP
| Entire | Name: Hexameric complex of TadA bound to ADP |
|---|---|
| Components |
|
-Supramolecule #1: Hexameric complex of TadA bound to ADP
| Supramolecule | Name: Hexameric complex of TadA bound to ADP / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Caulobacter vibrioides NA1000 (bacteria) Caulobacter vibrioides NA1000 (bacteria) |
-Macromolecule #1: Pilus assembly ATPase CpaF
| Macromolecule | Name: Pilus assembly ATPase CpaF / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Caulobacter vibrioides NA1000 (bacteria) Caulobacter vibrioides NA1000 (bacteria) |
| Molecular weight | Theoretical: 46.578355 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DYYHATKTTI FNALLNTIDL SQLAQLDLKQ AGEEIRDIVA ELVAIKNVSM SVAEQEHLVQ DIINDVLGYG PLEPLLARDD IADIMVNGA HRVFIEVGGK VQLTNVRFRD NLQLMNICQR IVSQVGRRVD ESSPICDARL PDGSRVNVIA PPLALDGPTL T IRKFKKDK ...String: DYYHATKTTI FNALLNTIDL SQLAQLDLKQ AGEEIRDIVA ELVAIKNVSM SVAEQEHLVQ DIINDVLGYG PLEPLLARDD IADIMVNGA HRVFIEVGGK VQLTNVRFRD NLQLMNICQR IVSQVGRRVD ESSPICDARL PDGSRVNVIA PPLALDGPTL T IRKFKKDK LTMKNLVEFA SISPEGARVL GVIGACRCNL VISGGTGSGK TTLLNTMTAF IDPTERVVTC EDAAELQLQQ PH VVRLETR PPNLEGSGAV TMRDLVKNCL RMRPERIIVG EVRGPEAFDL LQAMNTGHDG SMGTLHANSP REAISRIESM ITM GGYGLP SKTIKEMIVG SVDVIIQAAR LRDGSRRITH ITEVVGLEGD VIVTQDLFVY EITGEDEHGK VVGKHRSTGI ARPR FWDRA RYYGLERELA EALDAAEAL UniProtKB: Pilus assembly ATPase CpaF |
-Macromolecule #2: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 2 / Number of copies: 6 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Macromolecule #3: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 3 / Number of copies: 6 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #4: 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID
| Macromolecule | Name: 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID / type: ligand / ID: 4 / Number of copies: 4 / Formula: EPE |
|---|---|
| Molecular weight | Theoretical: 238.305 Da |
| Chemical component information |  ChemComp-EPE: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: INSILICO MODEL |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.1 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 114641 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller







 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)