[English] 日本語
 Yorodumi
Yorodumi- EMDB-18969: Structure of the SFTSV L protein stalled in a transcription-speci... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the SFTSV L protein stalled in a transcription-specific early elongation state with bound capped RNA [TRANSCRIPTION-EARLY-ELONGATION] | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | SFTSV / RNA-DEPENDENT RNA POLYMERASE / VIRAL RNA / VIRAL PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationvirion component / host cell endoplasmic reticulum-Golgi intermediate compartment / host cell Golgi apparatus / RNA-directed RNA polymerase / viral RNA genome replication / hydrolase activity / RNA-directed RNA polymerase activity / DNA-templated transcription / metal ion binding Similarity search - Function | ||||||||||||
| Biological species |  SFTS virus AH12 SFTS virus AH12 | ||||||||||||
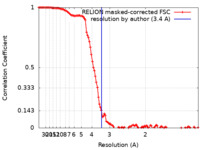
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | ||||||||||||
 Authors Authors | Williams HM / Thorkelsson SR / Vogel D / Busch C / Milewski M / Cusack S / Grunewald K / Quemin ERJ / Rosenthal M | ||||||||||||
| Funding support |  Germany, 3 items Germany, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2024 Journal: Nucleic Acids Res / Year: 2024Title: Structural snapshots of phenuivirus cap-snatching and transcription. Authors: Harry M Williams / Sigurdur R Thorkelsson / Dominik Vogel / Carola Busch / Morlin Milewski / Stephen Cusack / Kay Grünewald / Emmanuelle R J Quemin / Maria Rosenthal /   Abstract: Severe fever with thrombocytopenia syndrome virus (SFTSV) is a human pathogen that is now endemic to several East Asian countries. The viral large (L) protein catalyzes viral transcription by ...Severe fever with thrombocytopenia syndrome virus (SFTSV) is a human pathogen that is now endemic to several East Asian countries. The viral large (L) protein catalyzes viral transcription by stealing host mRNA caps via a process known as cap-snatching. Here, we establish an in vitro cap-snatching assay and present three high-quality electron cryo-microscopy (cryo-EM) structures of the SFTSV L protein in biologically relevant, transcription-specific states. In a priming-state structure, we show capped RNA bound to the L protein cap-binding domain (CBD). The L protein conformation in this priming structure is significantly different from published replication-state structures, in particular the N- and C-terminal domains. The capped-RNA is positioned in a way that it can feed directly into the RNA-dependent RNA polymerase (RdRp) ready for elongation. We also captured the L protein in an early-elongation state following primer-incorporation demonstrating that this priming conformation is retained at least in the very early stages of primer extension. This structural data is complemented by in vitro biochemical and cell-based assays. Together, these insights further our mechanistic understanding of how SFTSV and other bunyaviruses incorporate stolen host mRNA fragments into their viral transcripts thereby allowing the virus to hijack host cell translation machinery. #1:  Journal: BiorXiv / Year: 2023 Journal: BiorXiv / Year: 2023Title: Structural snapshots of phenuivirus cap-snatching and transcription Authors: Williams HM / Thorkelsson SR / Vogel D / Busch C / Milewski M / Cusack S / Grunewald K / Quemin ERJ / Rosenthal M | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18969.map.gz emd_18969.map.gz | 116.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18969-v30.xml emd-18969-v30.xml emd-18969.xml emd-18969.xml | 23.9 KB 23.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_18969_fsc.xml emd_18969_fsc.xml | 11.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_18969.png emd_18969.png | 103.4 KB | ||
| Filedesc metadata |  emd-18969.cif.gz emd-18969.cif.gz | 8.3 KB | ||
| Others |  emd_18969_half_map_1.map.gz emd_18969_half_map_1.map.gz emd_18969_half_map_2.map.gz emd_18969_half_map_2.map.gz | 98.4 MB 98.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18969 http://ftp.pdbj.org/pub/emdb/structures/EMD-18969 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18969 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18969 | HTTPS FTP |
-Related structure data
| Related structure data |  8r6yMC  8r6uC  8r6wC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_18969.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_18969.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.67 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_18969_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_18969_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : SFTSV L protein in complex with 5' and 3' RNA, as well as RNA act...
| Entire | Name: SFTSV L protein in complex with 5' and 3' RNA, as well as RNA acting as a primer. |
|---|---|
| Components |
|
-Supramolecule #1: SFTSV L protein in complex with 5' and 3' RNA, as well as RNA act...
| Supramolecule | Name: SFTSV L protein in complex with 5' and 3' RNA, as well as RNA acting as a primer. type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 Details: SFTSV L protein expressed recombinantly in complex with several RNA species. |
|---|---|
| Source (natural) | Organism:  SFTS virus AH12 SFTS virus AH12 |
| Molecular weight | Theoretical: 240 KDa |
-Macromolecule #1: RNA-directed RNA polymerase L
| Macromolecule | Name: RNA-directed RNA polymerase L / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  SFTS virus AH12 SFTS virus AH12 |
| Molecular weight | Theoretical: 235.6985 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MNLEVLCGRI NVENGLSLGE PGLYDQIYDR PGLPDLDVTV DATGVTVDIG AVPDSASQLG SSINAGLITI QLSEAYKINH DFTFSGLSK TTDRRLSEVF PITHDGSDGM TPAVIHTRLD GTIVVVEFST TRSHNIGGLE AAYRTKIEKY RDPISRRVDI M ENPRVFFG ...String: MNLEVLCGRI NVENGLSLGE PGLYDQIYDR PGLPDLDVTV DATGVTVDIG AVPDSASQLG SSINAGLITI QLSEAYKINH DFTFSGLSK TTDRRLSEVF PITHDGSDGM TPAVIHTRLD GTIVVVEFST TRSHNIGGLE AAYRTKIEKY RDPISRRVDI M ENPRVFFG VIVVSSGGVL SNMPLTQDEA EELMYRFCIA NEIYTKARSM DADIELQKSE EELEAISRAL SFFSLFEPNI ER VEGTFPN SEIKMLEQFL STPADVDFIT KTLKAKEVEA YADLCDSHYL KPEKTIQERL EINRCEAIDK TQDLLAGLHA RSN KQTSLN RGTVKLPPWL PKPSSESIDI KTDSGFGSLM DHGAYGELWA KCLLDVSLGN VEGVVSDPAK ELDIAISDDP EKDT PKEAK ITYRRFKPAL SSSARQEFSL QGVEGKKWKR MAANQKKEKE SHETLSPFLD VEDIGDFLTF NNLLTDSRYG DESIQ RAVS ILLEKASAMQ DTELTHALND SFKRNLSSNV VQWSLWVSCL AQELASALKQ HCRAGEFIIK KLKFWPIYVI IKPTKS SSH IFYSLGIRKA DVTRRLTGRV FSDTIDAGEW ELTEFKSLKT CKLTNLVNLP CTMLNSIAFW REKLGVAPWL VRKPCSE LR EQVGLTFLIS LEDKSKTEEI ITLTRYTQME GFVSPPMLPK PQKMLGKLDG PLRTKLQVYL LRKHLDCMVR IASQPFSL I PREGRVEWGG TFHAISGRST NLENMVNSWY IGYYKNKEES TELNALGEMY KKIVEMEEDK PSSPEFLGWG DTDSPKKHE FSRSFLRAAC SSLEREIAQR HGRQWKQNLE ERVLREIGTK NILDLASMKA TSNFSKDWEL YSEVQTKEYH RSKLLEKMAT LIEKGVMWY IDAVGQAWKA VLDDGCMRIC LFKKNQHGGL REIYVMDANA RLVQFGVETM ARCVCELSPH ETVANPRLKN S IIENHGLK SARSLGPGSI NINSSNDAKK WNQGHYTTKL ALVLCWFMPA KFHRFIWAAI SMFRRKKMMV DLRFLAHLSS KS ESRSSDP FREAMTDAFH GNRDVSWMDK GRTYIKTETG MMQGILHFTS SLLHSCVQSF YKSYFVSKLK EGYMGESISG VVD VIEGSD DSAIMISIRP KSDMDEVRSR FFVANLLHSV KFLNPLFGIY SSEKSTVNTV YCVEYNSEFH FHRHLVRPTL RWIA ASHQI SETEALASRQ EDYSNLLTQC LEGGASFSLT YLIQCAQLLH HYMLLGLCLH PLFGTFMGML ISDPDPALGF FLMDN PAFA GGAGFRFNLW RACKTTDLGR KYAYYFNEIQ GKTKGDEDYR ALDATSGGTL SHSVMVYWGD RKKYQALLNR MGLPED WVE QIDENPGVLY RRAANKKELL LKLAEKVHSP GVTSSLSKGH VVPRVVAAGV YLLSRHCFRF SSSIHGRGST QKASLIK LL MMSSISAMKH GGSLNPNQER MLFPQAQEYD RVCTLLEEVE HLTGKFVVRE RNIVRSRIDL FQEPVDLRCK AEDLVSEV W FGLKRTKLGP RLLKEEWDKL RASFAWLSTD PSETLRDGPF LSHVQFRNFI AHVDAKSRSV RLLGAPVKKS GGVTTISQV VRMNFFPGFS LEAEKSLDNQ ERLESISILK HVLFMVLNGP YTEEYKLEMI IEAFSTLVIP QPSEVIRKSR TMTLCLLSNY LSSRGGSIL DQIERAQSGT LGGFSKPQKT FVRPGGGVGY KGKGVWTGVM EDTHVQILID GDGTSNWLEE IRLSSDARLY D VIESIRRL CDDLGINNRV ASAYRGHCMV RLSGFKIKPA SRTDGCPVRI MERGFRIREL QNPDEVKMRV RGDILNLSVT IQ EGRVMNI LSYRPRDTDI SESAAAYLWS NRDLFSFGKK EPSCSWICLK TLDNWAWSHA SVLLANDRKT QGIDNRAMGN IFR DCLEGS LRKQGLMRSK LTEMVEKNVV PLTTQELVDI LEEDIDFSDV IAVELSEGSL DIESIFDGAP ILWSAEVEEF GEGV VAVSY SSKYYHLTLM DQAAITMCAI MGKEGCRGLL TEKRCMAAIR EQVRPFLIFL QIPEDSISWV SDQFCDSRGL DEEST IMWG UniProtKB: RNA-directed RNA polymerase L |
-Macromolecule #2: RNA (5'-R(*AP*CP*AP*CP*AP*GP*AP*GP*AP*CP*GP*CP*CP*CP*AP*G)-3')
| Macromolecule | Name: RNA (5'-R(*AP*CP*AP*CP*AP*GP*AP*GP*AP*CP*GP*CP*CP*CP*AP*G)-3') type: rna / ID: 2 Details: RNA corresponds to 5i end of SFTSV L genome segment. Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  SFTS virus AH12 SFTS virus AH12 |
| Molecular weight | Theoretical: 6.451975 KDa |
| Sequence | String: ACACAGAGAC GCCCAGAUGA |
-Macromolecule #3: RNA (5'-R(P*CP*UP*GP*GP*GP*CP*GP*GP*UP*AP*AP*AP*UP*GP*U)-3')
| Macromolecule | Name: RNA (5'-R(P*CP*UP*GP*GP*GP*CP*GP*GP*UP*AP*AP*AP*UP*GP*U)-3') type: rna / ID: 3 Details: RNA corresponds to 3' end of SFTSV L genome segment. Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  SFTS virus AH12 SFTS virus AH12 |
| Molecular weight | Theoretical: 6.480872 KDa |
| Sequence | String: GAUCUGGGCG GUAAAUGUGU |
-Macromolecule #4: RNA (5'-R(P*AP*CP*A)-3')
| Macromolecule | Name: RNA (5'-R(P*AP*CP*A)-3') / type: rna / ID: 4 / Details: Growing strand RNA. / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  SFTS virus AH12 SFTS virus AH12 |
| Molecular weight | Theoretical: 918.636 Da |
| Sequence | String: ACA |
-Macromolecule #5: RNA (5'-R(*(M7G)*AP*AP*A)-3')
| Macromolecule | Name: RNA (5'-R(*(M7G)*AP*AP*A)-3') / type: rna / ID: 5 Details: The ADM ligand is actually the m7GTP cap and the first A of the sequence. Hence, it corresponds to 2 nts *NOT* 1 nt. I have deleted the first 'real' A from the sequence to reflect this. Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  SFTS virus AH12 SFTS virus AH12 |
| Molecular weight | Theoretical: 5.863617 KDa |
| Sequence | String: (M7G)AAAAACGCA ACCAACAC |
-Macromolecule #6: 5'-O-[(S)-hydroxy{[(S)-hydroxy(phosphonooxy)phosphoryl]amino}phos...
| Macromolecule | Name: 5'-O-[(S)-hydroxy{[(S)-hydroxy(phosphonooxy)phosphoryl]amino}phosphoryl]uridine type: ligand / ID: 6 / Number of copies: 1 / Formula: 2KH |
|---|---|
| Molecular weight | Theoretical: 483.156 Da |
| Chemical component information | 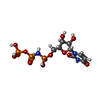 ChemComp-2KH: |
-Macromolecule #7: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 7 / Number of copies: 1 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.8 mg/mL |
|---|---|
| Buffer | pH: 7 |
| Grid | Model: Quantifoil R2/1 / Material: GOLD / Support film - Material: CARBON / Support film - topology: HOLEY |
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
| Details | This sample was monodisperse. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number real images: 27095 / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.6 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 130000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: RIGID BODY FIT / Target criteria: Cross-correlation coefficient |
|---|---|
| Output model |  PDB-8r6y: |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)