[English] 日本語
 Yorodumi
Yorodumi- EMDB-18592: E.coli DNA gyrase in complex with 217 bp substrate DNA and LEI-800 -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | E.coli DNA gyrase in complex with 217 bp substrate DNA and LEI-800 | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | TOPOISOMERASE / GYRASE / ALLOSTERIC INHIBITOR / ANTIBIOTIC / DNA BINDING PROTEIN | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationDNA negative supercoiling activity / DNA topoisomerase type II (double strand cut, ATP-hydrolyzing) complex / DNA topoisomerase type II (double strand cut, ATP-hydrolyzing) activity / DNA topoisomerase (ATP-hydrolysing) / DNA topological change / ATP-dependent activity, acting on DNA / DNA-templated DNA replication / chromosome / response to xenobiotic stimulus / response to antibiotic ...DNA negative supercoiling activity / DNA topoisomerase type II (double strand cut, ATP-hydrolyzing) complex / DNA topoisomerase type II (double strand cut, ATP-hydrolyzing) activity / DNA topoisomerase (ATP-hydrolysing) / DNA topological change / ATP-dependent activity, acting on DNA / DNA-templated DNA replication / chromosome / response to xenobiotic stimulus / response to antibiotic / DNA-templated transcription / DNA binding / ATP binding / metal ion binding / cytoplasm / cytosol Similarity search - Function | |||||||||||||||
| Biological species |   Escherichia phage Mu (virus) Escherichia phage Mu (virus) | |||||||||||||||
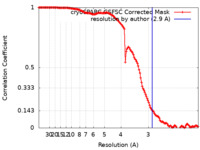
| Method | single particle reconstruction / cryo EM / Resolution: 2.9 Å | |||||||||||||||
 Authors Authors | Ghilarov D / Martin NI / van der Stelt M | |||||||||||||||
| Funding support |  United Kingdom, European Union, United Kingdom, European Union,  Netherlands, 4 items Netherlands, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Chem / Year: 2024 Journal: Nat Chem / Year: 2024Title: Discovery of isoquinoline sulfonamides as allosteric gyrase inhibitors with activity against fluoroquinolone-resistant bacteria. Authors: Alexander T Bakker / Ioli Kotsogianni / Mariana Avalos / Jeroen M Punt / Bing Liu / Diana Piermarini / Berend Gagestein / Cornelis J Slingerland / Le Zhang / Joost J Willemse / Leela B ...Authors: Alexander T Bakker / Ioli Kotsogianni / Mariana Avalos / Jeroen M Punt / Bing Liu / Diana Piermarini / Berend Gagestein / Cornelis J Slingerland / Le Zhang / Joost J Willemse / Leela B Ghimire / Richard J H B N van den Berg / Antonius P A Janssen / Tom H M Ottenhoff / Constant A A van Boeckel / Gilles P van Wezel / Dmitry Ghilarov / Nathaniel I Martin / Mario van der Stelt /   Abstract: Bacteria have evolved resistance to nearly all known antibacterials, emphasizing the need to identify antibiotics that operate via novel mechanisms. Here we report a class of allosteric inhibitors of ...Bacteria have evolved resistance to nearly all known antibacterials, emphasizing the need to identify antibiotics that operate via novel mechanisms. Here we report a class of allosteric inhibitors of DNA gyrase with antibacterial activity against fluoroquinolone-resistant clinical isolates of Escherichia coli. Screening of a small-molecule library revealed an initial isoquinoline sulfonamide hit, which was optimized via medicinal chemistry efforts to afford the more potent antibacterial LEI-800. Target identification studies, including whole-genome sequencing of in vitro selected mutants with resistance to isoquinoline sulfonamides, unanimously pointed to the DNA gyrase complex, an essential bacterial topoisomerase and an established antibacterial target. Using single-particle cryogenic electron microscopy, we determined the structure of the gyrase-LEI-800-DNA complex. The compound occupies an allosteric, hydrophobic pocket in the GyrA subunit and has a mode of action that is distinct from the clinically used fluoroquinolones or any other gyrase inhibitor reported to date. LEI-800 provides a chemotype suitable for development to counter the increasingly widespread bacterial resistance to fluoroquinolones. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18592.map.gz emd_18592.map.gz | 230.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18592-v30.xml emd-18592-v30.xml emd-18592.xml emd-18592.xml | 23.2 KB 23.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_18592_fsc.xml emd_18592_fsc.xml | 13.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_18592.png emd_18592.png | 47.9 KB | ||
| Filedesc metadata |  emd-18592.cif.gz emd-18592.cif.gz | 7.5 KB | ||
| Others |  emd_18592_additional_1.map.gz emd_18592_additional_1.map.gz emd_18592_half_map_1.map.gz emd_18592_half_map_1.map.gz emd_18592_half_map_2.map.gz emd_18592_half_map_2.map.gz | 204.7 MB 226.3 MB 226.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18592 http://ftp.pdbj.org/pub/emdb/structures/EMD-18592 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18592 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18592 | HTTPS FTP |
-Validation report
| Summary document |  emd_18592_validation.pdf.gz emd_18592_validation.pdf.gz | 985.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_18592_full_validation.pdf.gz emd_18592_full_validation.pdf.gz | 985.5 KB | Display | |
| Data in XML |  emd_18592_validation.xml.gz emd_18592_validation.xml.gz | 22.2 KB | Display | |
| Data in CIF |  emd_18592_validation.cif.gz emd_18592_validation.cif.gz | 28.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18592 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18592 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18592 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18592 | HTTPS FTP |
-Related structure data
| Related structure data |  8qqiMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_18592.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_18592.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.02 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #1
| File | emd_18592_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_18592_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_18592_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Ternary complex of E.coli DNA gyrase with 217 bp linear dsDNA fra...
| Entire | Name: Ternary complex of E.coli DNA gyrase with 217 bp linear dsDNA fragment and LEI-800 |
|---|---|
| Components |
|
-Supramolecule #1: Ternary complex of E.coli DNA gyrase with 217 bp linear dsDNA fra...
| Supramolecule | Name: Ternary complex of E.coli DNA gyrase with 217 bp linear dsDNA fragment and LEI-800 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: DNA gyrase subunit A
| Macromolecule | Name: DNA gyrase subunit A / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 58.824207 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SSDLAREITP VNIEEELKSS YLDYAMSVIV GRALPDVRDG LKPVHRRVLY AMNVLGNDWN KAYKKSARVV GDVIGKYHPH GDSAVYDTI VRMAQPFSLR YMLVDGQGNF GSIDGDSAAA MRYTEIRLAK IAHELMADLE KETVDFVDNY DGTEKIPDVM P TKIPNLLV ...String: SSDLAREITP VNIEEELKSS YLDYAMSVIV GRALPDVRDG LKPVHRRVLY AMNVLGNDWN KAYKKSARVV GDVIGKYHPH GDSAVYDTI VRMAQPFSLR YMLVDGQGNF GSIDGDSAAA MRYTEIRLAK IAHELMADLE KETVDFVDNY DGTEKIPDVM P TKIPNLLV NGSSGIAVGM ATNIPPHNLT EVINGCLAYI DDEDISIEGL MEHIPGPDFP TAAIINGRRG IEEAYRTGRG KV YIRARAE VEVDAKTGRE TIIVHEIPYQ VNKARLIEKI AELVKEKRVE GISALRDESD KDGMRIVIEV KRDAVGEVVL NNL YSQTQL QVSFGINMVA LHHGQPKIMN LKDIIAAFVR HRREVVTRRT IFELRKARDR AHILEALAVA LANIDPIIEL IRHA PTPAE AKTALVANPW QLGNVAAMLE RAGDDAARPE WLEPEFGVRD GLYYLTEQQA QAILDLRLQK LTGLEHEKLL DEYKE LLDQ IAELLRILGS ADRLMEVIRE ELELVREQFG DKRRTEITAN UniProtKB: DNA gyrase subunit A |
-Macromolecule #2: DNA gyrase subunit B
| Macromolecule | Name: DNA gyrase subunit B / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO / EC number: DNA topoisomerase (ATP-hydrolysing) |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 89.641391 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GPSNSYDSSS IKVLKGLDAV RKRPGMYIGD TDDGTGLHHM VFEVVDNAID EALAGHCKEI IVTIHADNSV SVQDDGRGIP TGIHPEEGV SAAEVIMTVL HAGGKFDDNS YKVSGGLHGV GVSVVNALSQ KLELVIQREG KIHRQIYEHG VPQAPLAVTG E TEKTGTMV ...String: GPSNSYDSSS IKVLKGLDAV RKRPGMYIGD TDDGTGLHHM VFEVVDNAID EALAGHCKEI IVTIHADNSV SVQDDGRGIP TGIHPEEGV SAAEVIMTVL HAGGKFDDNS YKVSGGLHGV GVSVVNALSQ KLELVIQREG KIHRQIYEHG VPQAPLAVTG E TEKTGTMV RFWPSLETFT NVTEFEYEIL AKRLRELSFL NSGVSIRLRD KRDGKEDHFH YEGGIKAFVE YLNKNKTPIH PN IFYFSTE KDGIGVEVAL QWNDGFQENI YCFTNNIPQR DGGTHLAGFR AAMTRTLNAY MDKEGYSKKA KVSATGDDAR EGL IAVVSV KVPDPKFSSQ TKDKLVSSEV KSAVEQQMNE LLAEYLLENP TDAKIVVGKI IDAARAREAA RRAREMTRRK GALD LAGLP GKLADCQERD PALSELYLVE GDSAGGSAKQ GRNRKNQAIL PLKGKILNVE KARFDKMLSS QEVATLITAL GCGIG RDEY NPDKLRYHSI IIMTDADVDG SHIRTLLLTF FYRQMPEIVE RGHVYIAQPP LYKVKKGKQE QYIKDDEAMD QYQISI ALD GATLHTNASA PALAGEALEK LVSEYNATQK MINRMERRYP KAMLKELIYQ PTLTEADLSD EQTVTRWVNA LVSELND KE QHGSQWKFDV HTNAEQNLFE PIVRVRTHGV DTDYPLDHEF ITGGEYRRIC TLGEKLRGLL EEDAFIERGE RRQPVASF E QALDWLVKES RRGLSIQRYK GLGEMNPEQL WETTMDPESR RMLRVTVKDA IAADQLFTTL MGDAVEPRRA FIEENALKA A UniProtKB: DNA gyrase subunit B |
-Macromolecule #3: Mu217 chain E
| Macromolecule | Name: Mu217 chain E / type: dna / ID: 3 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Escherichia phage Mu (virus) Escherichia phage Mu (virus) |
| Molecular weight | Theoretical: 7.943173 KDa |
| Sequence | String: (DA)(DT)(DC)(DA)(DG)(DG)(DC)(DA)(DT)(DA) (DA)(DA)(DA)(DT)(DC)(DA)(DC)(DC)(DC)(DG) (DC)(DA)(DC)(DA)(DG)(DA) |
-Macromolecule #4: Mu217 chain F
| Macromolecule | Name: Mu217 chain F / type: dna / ID: 4 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Escherichia phage Mu (virus) Escherichia phage Mu (virus) |
| Molecular weight | Theoretical: 8.031157 KDa |
| Sequence | String: (DT)(DC)(DT)(DG)(DT)(DG)(DC)(DG)(DG)(DG) (DT)(DG)(DA)(DT)(DT)(DT)(DT)(DA)(DT)(DG) (DC)(DC)(DT)(DG)(DA)(DT) |
-Macromolecule #5: ~{N}-[[(2~{S},5~{R})-5-(4-pyridin-3-ylphenyl)pyrrolidin-2-yl]meth...
| Macromolecule | Name: ~{N}-[[(2~{S},5~{R})-5-(4-pyridin-3-ylphenyl)pyrrolidin-2-yl]methyl]isoquinoline-5-sulfonamide type: ligand / ID: 5 / Number of copies: 2 / Formula: LRL |
|---|---|
| Molecular weight | Theoretical: 444.549 Da |
-Macromolecule #6: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 6 / Number of copies: 2 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 12 mg/mL |
|---|---|
| Buffer | pH: 8 Details: 25 mM Na-HEPES pH 8 KOAc 30 mM MgOAc 2.5 mM TCEP 0.5 mM CHAPSO 8 mM |
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 300 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 293 K / Instrument: FEI VITROBOT MARK IV |
| Details | Monodisperse sample |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 39.58 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.8000000000000003 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






 Z
Z Y
Y X
X