[English] 日本語
 Yorodumi
Yorodumi- EMDB-18460: mt-LSU assembly intermediate in GTPBP8 knock-out cells, state 1 -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | mt-LSU assembly intermediate in GTPBP8 knock-out cells, state 1 | |||||||||
 Map data Map data | mt-LSU assembly intermediate in GTPBP8 knock-out cells, state 1 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Mitochondria / Assembly / GTPBP8 / RIBOSOME | |||||||||
| Function / homology |  Function and homology information Function and homology informationrRNA import into mitochondrion / Mitochondrial ribosome-associated quality control / Mitochondrial translation elongation / Mitochondrial translation initiation / Mitochondrial translation termination / mitochondrial large ribosomal subunit / mitochondrial ribosome / mitochondrial translation / Mitochondrial protein degradation / cellular response to leukemia inhibitory factor ...rRNA import into mitochondrion / Mitochondrial ribosome-associated quality control / Mitochondrial translation elongation / Mitochondrial translation initiation / Mitochondrial translation termination / mitochondrial large ribosomal subunit / mitochondrial ribosome / mitochondrial translation / Mitochondrial protein degradation / cellular response to leukemia inhibitory factor / fibrillar center / 5S rRNA binding / large ribosomal subunit rRNA binding / mitochondrial inner membrane / negative regulation of translation / rRNA binding / structural constituent of ribosome / ribosome / translation / mitochondrial matrix / ribonucleoprotein complex / protein domain specific binding / mRNA binding / mitochondrion / extracellular space / RNA binding / nucleoplasm / nucleus / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
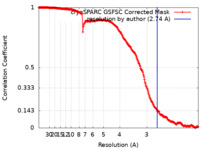
| Method | single particle reconstruction / cryo EM / Resolution: 2.74 Å | |||||||||
 Authors Authors | Valentin Gese G / Cipullo M / Rorbach J / Hallberg BM | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: GTPBP8 plays a role in mitoribosome formation in human mitochondria. Authors: Miriam Cipullo / Genís Valentín Gesé / Shreekara Gopalakrishna / Annika Krueger / Vivian Lobo / Maria A Pirozhkova / James Marks / Petra Páleníková / Dmitrii Shiriaev / Yong Liu / ...Authors: Miriam Cipullo / Genís Valentín Gesé / Shreekara Gopalakrishna / Annika Krueger / Vivian Lobo / Maria A Pirozhkova / James Marks / Petra Páleníková / Dmitrii Shiriaev / Yong Liu / Jelena Misic / Yu Cai / Minh Duc Nguyen / Abubakar Abdelbagi / Xinping Li / Michal Minczuk / Markus Hafner / Daniel Benhalevy / Aishe A Sarshad / Ilian Atanassov / B Martin Hällberg / Joanna Rorbach /      Abstract: Mitochondrial gene expression relies on mitoribosomes to translate mitochondrial mRNAs. The biogenesis of mitoribosomes is an intricate process involving multiple assembly factors. Among these ...Mitochondrial gene expression relies on mitoribosomes to translate mitochondrial mRNAs. The biogenesis of mitoribosomes is an intricate process involving multiple assembly factors. Among these factors, GTP-binding proteins (GTPBPs) play important roles. In bacterial systems, numerous GTPBPs are required for ribosome subunit maturation, with EngB being a GTPBP involved in the ribosomal large subunit assembly. In this study, we focus on exploring the function of GTPBP8, the human homolog of EngB. We find that ablation of GTPBP8 leads to the inhibition of mitochondrial translation, resulting in significant impairment of oxidative phosphorylation. Structural analysis of mitoribosomes from GTPBP8 knock-out cells shows the accumulation of mitoribosomal large subunit assembly intermediates that are incapable of forming functional monosomes. Furthermore, fPAR-CLIP analysis reveals that GTPBP8 is an RNA-binding protein that interacts specifically with the mitochondrial ribosome large subunit 16 S rRNA. Our study highlights the role of GTPBP8 as a component of the mitochondrial gene expression machinery involved in mitochondrial large subunit maturation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18460.map.gz emd_18460.map.gz | 775.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18460-v30.xml emd-18460-v30.xml emd-18460.xml emd-18460.xml | 14.5 KB 14.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_18460_fsc.xml emd_18460_fsc.xml | 19.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_18460.png emd_18460.png | 58.9 KB | ||
| Masks |  emd_18460_msk_1.map emd_18460_msk_1.map | 824 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-18460.cif.gz emd-18460.cif.gz | 4.2 KB | ||
| Others |  emd_18460_half_map_1.map.gz emd_18460_half_map_1.map.gz emd_18460_half_map_2.map.gz emd_18460_half_map_2.map.gz | 763.4 MB 763.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18460 http://ftp.pdbj.org/pub/emdb/structures/EMD-18460 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18460 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18460 | HTTPS FTP |
-Related structure data
| Related structure data |  8qu1MC  8qrkC  8qrlC  8qrmC  8qrnC  8qu5C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_18460.map.gz / Format: CCP4 / Size: 824 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_18460.map.gz / Format: CCP4 / Size: 824 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | mt-LSU assembly intermediate in GTPBP8 knock-out cells, state 1 | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.01 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_18460_msk_1.map emd_18460_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: mt-LSU assembly intermediate in GTPBP8 knock-out cells, state...
| File | emd_18460_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | mt-LSU assembly intermediate in GTPBP8 knock-out cells, state 1, half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: mt-LSU assembly intermediate in GTPBP8 knock-out cells, state...
| File | emd_18460_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | mt-LSU assembly intermediate in GTPBP8 knock-out cells, state 1, half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : mt-SSU assembly intermediate in GTPBP8 knock-out cells, state 2
| Entire | Name: mt-SSU assembly intermediate in GTPBP8 knock-out cells, state 2 |
|---|---|
| Components |
|
-Supramolecule #1: mt-SSU assembly intermediate in GTPBP8 knock-out cells, state 2
| Supramolecule | Name: mt-SSU assembly intermediate in GTPBP8 knock-out cells, state 2 type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 1.7 MDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 48.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)