+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Chimeric Adenovirus-derived dodecamer | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Adenovirus / capsid protein / virus-like particle / VIRUS LIKE PARTICLE | |||||||||
| Function / homology | Adenovirus penton base protein / Adenovirus penton base protein / T=25 icosahedral viral capsid / endocytosis involved in viral entry into host cell / virion attachment to host cell / host cell nucleus / structural molecule activity / metal ion binding / Penton protein Function and homology information Function and homology information | |||||||||
| Biological species |  Human adenovirus sp. / Human adenovirus sp. /  unidentified adenovirus unidentified adenovirus | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.2 Å | |||||||||
 Authors Authors | Buzas D / Borucu U / Bufton J / Kapadalakere SY / Toelzer C | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Structure / Year: 2024 Journal: Structure / Year: 2024Title: Engineering the ADDobody protein scaffold for generation of high-avidity ADDomer super-binders. Authors: Dora Buzas / Huan Sun / Christine Toelzer / Sathish K N Yadav / Ufuk Borucu / Gunjan Gautam / Kapil Gupta / Joshua C Bufton / Julien Capin / Richard B Sessions / Frederic Garzoni / Imre ...Authors: Dora Buzas / Huan Sun / Christine Toelzer / Sathish K N Yadav / Ufuk Borucu / Gunjan Gautam / Kapil Gupta / Joshua C Bufton / Julien Capin / Richard B Sessions / Frederic Garzoni / Imre Berger / Christiane Schaffitzel /  Abstract: Adenovirus-derived nanoparticles (ADDomer) comprise 60 copies of adenovirus penton base protein (PBP). ADDomer is thermostable, rendering the storage, transport, and deployment of ADDomer-based ...Adenovirus-derived nanoparticles (ADDomer) comprise 60 copies of adenovirus penton base protein (PBP). ADDomer is thermostable, rendering the storage, transport, and deployment of ADDomer-based therapeutics independent of a cold chain. To expand the scope of ADDomers for new applications, we engineered ADDobodies, representing PBP crown domain, genetically separated from PBP multimerization domain. We inserted heterologous sequences into hyper-variable loops, resulting in monomeric, thermostable ADDobodies expressed at high yields in Escherichia coli. The X-ray structure of an ADDobody prototype validated our design. ADDobodies can be used in ribosome display experiments to select a specific binder against a target, with an enrichment factor of ∼10-fold per round. ADDobodies can be re-converted into ADDomers by genetically reconnecting the selected ADDobody with the PBP multimerization domain from a different species, giving rise to a multivalent nanoparticle, called Chimera, confirmed by a 2.2 Å electron cryo-microscopy structure. Chimera comprises 60 binding sites, resulting in ultra-high, picomolar avidity to the target. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18323.map.gz emd_18323.map.gz | 36.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18323-v30.xml emd-18323-v30.xml emd-18323.xml emd-18323.xml | 14.3 KB 14.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_18323_fsc.xml emd_18323_fsc.xml | 14 KB | Display |  FSC data file FSC data file |
| Images |  emd_18323.png emd_18323.png | 77.2 KB | ||
| Masks |  emd_18323_msk_1.map emd_18323_msk_1.map | 244.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-18323.cif.gz emd-18323.cif.gz | 5.7 KB | ||
| Others |  emd_18323_half_map_1.map.gz emd_18323_half_map_1.map.gz emd_18323_half_map_2.map.gz emd_18323_half_map_2.map.gz | 190.8 MB 190.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18323 http://ftp.pdbj.org/pub/emdb/structures/EMD-18323 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18323 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18323 | HTTPS FTP |
-Related structure data
| Related structure data |  8qbxMC  8coiC  8qb3C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_18323.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_18323.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.05 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_18323_msk_1.map emd_18323_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
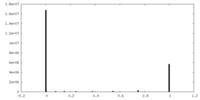
| Density Histograms |
-Half map: half-map1
| File | emd_18323_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half-map1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half-map2
| File | emd_18323_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half-map2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : unidentified adenovirus
| Entire | Name:  unidentified adenovirus unidentified adenovirus |
|---|---|
| Components |
|
-Supramolecule #1: unidentified adenovirus
| Supramolecule | Name: unidentified adenovirus / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 10535 / Sci species name: unidentified adenovirus / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: OTHER / Virus enveloped: No / Virus empty: Yes |
|---|
-Macromolecule #1: Penton protein
| Macromolecule | Name: Penton protein / type: protein_or_peptide / ID: 1 / Number of copies: 60 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Human adenovirus sp. Human adenovirus sp. |
| Molecular weight | Theoretical: 63.195812 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MMRRAYPEGP PPSYESVMQQ AMAAAAAMQP PLEAPYVPPR YLAPTEGRNS IRYSELSPLY DTTRLYLVDN KSADIASLNY QNDHSNFLT TVVQNNDFTP TEASTQTINF DERSRWGGQL KTIMHTNMPN VNEYMFSNKF KARVMVSRKA PEGEFVTVND G PVNDTYDH ...String: MMRRAYPEGP PPSYESVMQQ AMAAAAAMQP PLEAPYVPPR YLAPTEGRNS IRYSELSPLY DTTRLYLVDN KSADIASLNY QNDHSNFLT TVVQNNDFTP TEASTQTINF DERSRWGGQL KTIMHTNMPN VNEYMFSNKF KARVMVSRKA PEGEFVTVND G PVNDTYDH KEDILKYEWF EFILPEGNFS ATMTIDLMNN AIIDNYLEIG RQNGVLESDI GVKFDTRNFR LGWDPETKLI MP GVYTYEA FHPDIVLLPG CGVDFTESRL SNLLGIRKRH PFQEGFKIMY EDLEGGNIPA LLDVTAYEES KKDTTTARET TTL AVAEET SEDVDDDITR GDTYITELEK QKREAAAAEV SRKKELKIQP LEKDSKSRSY NVLEDKINTA YRSWYLSYNY GNPE KGIRS WTLLTTSDVT CGVEQVYWSL PDMMQDPVTF RSTRQVSNYP VVGAELMPVF SKSFYNEQAV YSQQLRQATS LTHVF NRFP ENQILIRPPA PTITTVSENV PALTDHGTLP LRSSIRGVQR VTVTDARRRT CPYVYKALGI VAPRVLSSRT F UniProtKB: Penton protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 1.06 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)