[English] 日本語
 Yorodumi
Yorodumi- EMDB-18135: Cryo-EM structure of the methanogenic Na+ translocating N5-methyl... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the methanogenic Na+ translocating N5-methyl-H4MPT:CoM methyltransferase complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | methanogenesis / tetrahydromethanopterin / coenzyme M / vitamin B12 / Na+ transport / TRANSFERASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationtetrahydromethanopterin S-methyltransferase / tetrahydromethanopterin S-methyltransferase activity / methyltransferase complex / methanogenesis, from carbon dioxide / vesicle membrane / cobalt ion binding / one-carbon metabolic process / methylation / plasma membrane / cytoplasm Similarity search - Function | |||||||||
| Biological species |   Methanothermobacter marburgensis (archaea) Methanothermobacter marburgensis (archaea) | |||||||||
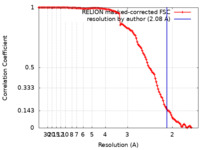
| Method | single particle reconstruction / cryo EM / Resolution: 2.08 Å | |||||||||
 Authors Authors | Aziz I / Vonck J / Ermler U | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2024 Journal: Proc Natl Acad Sci U S A / Year: 2024Title: Structural and mechanistic basis of the central energy-converting methyltransferase complex of methanogenesis. Authors: Iram Aziz / Kanwal Kayastha / Susann Kaltwasser / Janet Vonck / Sonja Welsch / Bonnie J Murphy / Jörg Kahnt / Di Wu / Tristan Wagner / Seigo Shima / Ulrich Ermler /  Abstract: Methanogenic archaea inhabiting anaerobic environments play a crucial role in the global biogeochemical material cycle. The most universal electrogenic reaction of their methane-producing energy ...Methanogenic archaea inhabiting anaerobic environments play a crucial role in the global biogeochemical material cycle. The most universal electrogenic reaction of their methane-producing energy metabolism is catalyzed by -methyl-tetrahydromethanopterin: coenzyme M methyltransferase (MtrABCDEFGH), which couples the vectorial Na transport with a methyl transfer between the one-carbon carriers tetrahydromethanopterin and coenzyme M via a vitamin B derivative (cobamide) as prosthetic group. We present the 2.08 Å cryo-EM structure of Mtr(ABCDEFG) composed of the central Mtr(ABFG) stalk symmetrically flanked by three membrane-spanning MtrCDE globes. Tetraether glycolipids visible in the map fill gaps inside the multisubunit complex. Putative coenzyme M and Na were identified inside or in a side-pocket of a cytoplasmic cavity formed within MtrCDE. Its bottom marks the gate of the transmembrane pore occluded in the cryo-EM map. By integrating Alphafold2 information, functionally competent MtrA-MtrH and MtrA-MtrCDE subcomplexes could be modeled and thus the methyl-tetrahydromethanopterin demethylation and coenzyme M methylation half-reactions structurally described. Methyl-transfer-driven Na transport is proposed to be based on a strong and weak complex between MtrCDE and MtrA carrying vitamin B, the latter being placed at the entrance of the cytoplasmic MtrCDE cavity. Hypothetically, strongly attached methyl-cob(III)amide (His-on) carrying MtrA induces an inward-facing conformation, Na flux into the membrane protein center and finally coenzyme M methylation while the generated loosely attached (or detached) MtrA carrying cob(I)amide (His-off) induces an outward-facing conformation and an extracellular Na outflux. Methyl-cob(III)amide (His-on) is regenerated in the distant active site of the methyl-tetrahydromethanopterin binding MtrH implicating a large-scale shuttling movement of the vitamin B-carrying domain. | |||||||||
| History |
|
- Structure visualization
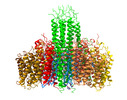
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18135.map.gz emd_18135.map.gz | 98.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18135-v30.xml emd-18135-v30.xml emd-18135.xml emd-18135.xml | 21.9 KB 21.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_18135_fsc.xml emd_18135_fsc.xml | 11.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_18135.png emd_18135.png | 119.9 KB | ||
| Masks |  emd_18135_msk_1.map emd_18135_msk_1.map | 125 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-18135.cif.gz emd-18135.cif.gz | 6.6 KB | ||
| Others |  emd_18135_half_map_1.map.gz emd_18135_half_map_1.map.gz emd_18135_half_map_2.map.gz emd_18135_half_map_2.map.gz | 98.4 MB 98.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18135 http://ftp.pdbj.org/pub/emdb/structures/EMD-18135 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18135 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18135 | HTTPS FTP |
-Related structure data
| Related structure data |  8q3vMC  8q54C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_18135.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_18135.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.837 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_18135_msk_1.map emd_18135_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
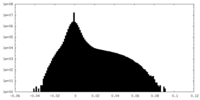
| Density Histograms |
-Half map: #2
| File | emd_18135_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_18135_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Methyl-H4MPT:CoM methyltransferase
+Supramolecule #1: Methyl-H4MPT:CoM methyltransferase
+Macromolecule #1: Tetrahydromethanopterin S-methyltransferase subunit A 1
+Macromolecule #2: Tetrahydromethanopterin S-methyltransferase subunit B
+Macromolecule #3: Tetrahydromethanopterin S-methyltransferase subunit C
+Macromolecule #4: Tetrahydromethanopterin S-methyltransferase subunit D
+Macromolecule #5: Tetrahydromethanopterin S-methyltransferase subunit E
+Macromolecule #6: Tetrahydromethanopterin S-methyltransferase subunit F
+Macromolecule #7: Tetrahydromethanopterin S-methyltransferase subunit G
+Macromolecule #8: MAGNESIUM ION
+Macromolecule #9: [(2~{S},7~{R},11~{R},15~{S},19~{S},22~{S},26~{S},30~{R},34~{R},39...
+Macromolecule #10: SODIUM ION
+Macromolecule #11: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 73.9 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.1 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)