[English] 日本語
 Yorodumi
Yorodumi- EMDB-18126: Structural studies of human serum albumin using cryo-EM up to 0.3... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structural studies of human serum albumin using cryo-EM up to 0.38 nm resolution | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | human serum albumin / monomer / TRANSPORT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationCiprofloxacin ADME / exogenous protein binding / cellular response to calcium ion starvation / enterobactin binding / Heme biosynthesis / HDL remodeling / negative regulation of mitochondrial depolarization / Prednisone ADME / Heme degradation / Aspirin ADME ...Ciprofloxacin ADME / exogenous protein binding / cellular response to calcium ion starvation / enterobactin binding / Heme biosynthesis / HDL remodeling / negative regulation of mitochondrial depolarization / Prednisone ADME / Heme degradation / Aspirin ADME / antioxidant activity / toxic substance binding / Scavenging of heme from plasma / Recycling of bile acids and salts / platelet alpha granule lumen / fatty acid binding / cellular response to starvation / Post-translational protein phosphorylation / Cytoprotection by HMOX1 / Regulation of Insulin-like Growth Factor (IGF) transport and uptake by Insulin-like Growth Factor Binding Proteins (IGFBPs) / pyridoxal phosphate binding / Platelet degranulation / protein-folding chaperone binding / blood microparticle / endoplasmic reticulum lumen / copper ion binding / endoplasmic reticulum / Golgi apparatus / protein-containing complex / extracellular space / DNA binding / extracellular exosome / extracellular region / identical protein binding / nucleus / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.77 Å | |||||||||
 Authors Authors | Slawek J / Taube M / Rawski M / Wojciechowska D / Kozak M | |||||||||
| Funding support |  Poland, 1 items Poland, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Structural studies of human serum albumin using cryo-EM up to 0.38 nm resolution Authors: Slawek J / Taube M / Rawski M / Wojciechowska D / Kozak M | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18126.map.gz emd_18126.map.gz | 31.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18126-v30.xml emd-18126-v30.xml emd-18126.xml emd-18126.xml | 16.2 KB 16.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_18126.png emd_18126.png | 44.5 KB | ||
| Filedesc metadata |  emd-18126.cif.gz emd-18126.cif.gz | 6 KB | ||
| Others |  emd_18126_half_map_1.map.gz emd_18126_half_map_1.map.gz emd_18126_half_map_2.map.gz emd_18126_half_map_2.map.gz | 59.4 MB 59.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18126 http://ftp.pdbj.org/pub/emdb/structures/EMD-18126 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18126 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18126 | HTTPS FTP |
-Validation report
| Summary document |  emd_18126_validation.pdf.gz emd_18126_validation.pdf.gz | 743.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_18126_full_validation.pdf.gz emd_18126_full_validation.pdf.gz | 743.1 KB | Display | |
| Data in XML |  emd_18126_validation.xml.gz emd_18126_validation.xml.gz | 12 KB | Display | |
| Data in CIF |  emd_18126_validation.cif.gz emd_18126_validation.cif.gz | 14.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18126 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18126 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18126 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18126 | HTTPS FTP |
-Related structure data
| Related structure data |  8q3fMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_18126.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_18126.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.86 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_18126_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
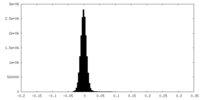
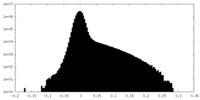
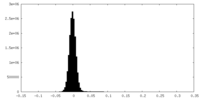
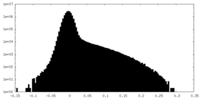
| Density Histograms |
-Half map: #1
| File | emd_18126_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Monomer of human serum albumin
| Entire | Name: Monomer of human serum albumin |
|---|---|
| Components |
|
-Supramolecule #1: Monomer of human serum albumin
| Supramolecule | Name: Monomer of human serum albumin / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 67 KDa |
-Macromolecule #1: Serum albumin
| Macromolecule | Name: Serum albumin / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 66.571219 KDa |
| Sequence | String: DAHKSEVAHR FKDLGEENFK ALVLIAFAQY LQQCPFEDHV KLVNEVTEFA KTCVADESAE NCDKSLHTLF GDKLCTVATL RETYGEMAD CCAKQEPERN ECFLQHKDDN PNLPRLVRPE VDVMCTAFHD NEETFLKKYL YEIARRHPYF YAPELLFFAK R YKAAFTEC ...String: DAHKSEVAHR FKDLGEENFK ALVLIAFAQY LQQCPFEDHV KLVNEVTEFA KTCVADESAE NCDKSLHTLF GDKLCTVATL RETYGEMAD CCAKQEPERN ECFLQHKDDN PNLPRLVRPE VDVMCTAFHD NEETFLKKYL YEIARRHPYF YAPELLFFAK R YKAAFTEC CQAADKAACL LPKLDELRDE GKASSAKQRL KCASLQKFGE RAFKAWAVAR LSQRFPKAEF AEVSKLVTDL TK VHTECCH GDLLECADDR ADLAKYICEN QDSISSKLKE CCEKPLLEKS HCIAEVENDE MPADLPSLAA DFVESKDVCK NYA EAKDVF LGMFLYEYAR RHPDYSVVLL LRLAKTYETT LEKCCAAADP HECYAKVFDE FKPLVEEPQN LIKQNCELFE QLGE YKFQN ALLVRYTKKV PQVSTPTLVE VSRNLGKVGS KCCKHPEAKR MPCAEDYLSV VLNQLCVLHE KTPVSDRVTK CCTES LVNR RPCFSALEVD ETYVPKEFNA ETFTFHADIC TLSEKERQIK KQTALVELVK HKPKATKEQL KAVMDDFAAF VEKCCK ADD KETCFAEEGK KLVAASQAAL GL UniProtKB: Albumin |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 Details: PBS buffer at pH=7.4 (137 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, 1.8 mM KH2PO4) |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Number real images: 11116 / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.9 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)