+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Murine type III Abeta fibril from ARTE10 mouse | ||||||||||||||||||
 Map data Map data | Ex vivo Abeta40 fibril from ARTE10 mouse brain. | ||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | Amyloid fibril / PROTEIN FIBRIL | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationcytosolic mRNA polyadenylation / collateral sprouting in absence of injury / microglia development / regulation of Wnt signaling pathway / regulation of synapse structure or activity / Formyl peptide receptors bind formyl peptides and many other ligands / axo-dendritic transport / synaptic assembly at neuromuscular junction / signaling receptor activator activity / axon midline choice point recognition ...cytosolic mRNA polyadenylation / collateral sprouting in absence of injury / microglia development / regulation of Wnt signaling pathway / regulation of synapse structure or activity / Formyl peptide receptors bind formyl peptides and many other ligands / axo-dendritic transport / synaptic assembly at neuromuscular junction / signaling receptor activator activity / axon midline choice point recognition / smooth endoplasmic reticulum calcium ion homeostasis / astrocyte activation involved in immune response / regulation of spontaneous synaptic transmission / mating behavior / NMDA selective glutamate receptor signaling pathway / ciliary rootlet / Lysosome Vesicle Biogenesis / PTB domain binding / Golgi-associated vesicle / Insertion of tail-anchored proteins into the endoplasmic reticulum membrane / positive regulation of amyloid fibril formation / neuron remodeling / Deregulated CDK5 triggers multiple neurodegenerative pathways in Alzheimer's disease models / nuclear envelope lumen / suckling behavior / COPII-coated ER to Golgi transport vesicle / dendrite development / presynaptic active zone / modulation of excitatory postsynaptic potential / TRAF6 mediated NF-kB activation / Advanced glycosylation endproduct receptor signaling / neuromuscular process controlling balance / The NLRP3 inflammasome / negative regulation of long-term synaptic potentiation / regulation of presynapse assembly / transition metal ion binding / regulation of multicellular organism growth / negative regulation of neuron differentiation / intracellular copper ion homeostasis / ECM proteoglycans / spindle midzone / positive regulation of T cell migration / smooth endoplasmic reticulum / Purinergic signaling in leishmaniasis infection / protein serine/threonine kinase binding / regulation of peptidyl-tyrosine phosphorylation / clathrin-coated pit / positive regulation of chemokine production / forebrain development / Notch signaling pathway / neuron projection maintenance / positive regulation of G2/M transition of mitotic cell cycle / Mitochondrial protein degradation / positive regulation of protein metabolic process / positive regulation of calcium-mediated signaling / ionotropic glutamate receptor signaling pathway / cholesterol metabolic process / positive regulation of glycolytic process / extracellular matrix organization / positive regulation of mitotic cell cycle / response to interleukin-1 / axonogenesis / adult locomotory behavior / trans-Golgi network membrane / platelet alpha granule lumen / positive regulation of peptidyl-threonine phosphorylation / dendritic shaft / learning / positive regulation of interleukin-1 beta production / positive regulation of long-term synaptic potentiation / central nervous system development / endosome lumen / locomotory behavior / astrocyte activation / Post-translational protein phosphorylation / positive regulation of JNK cascade / microglial cell activation / synapse organization / regulation of long-term neuronal synaptic plasticity / TAK1-dependent IKK and NF-kappa-B activation / serine-type endopeptidase inhibitor activity / visual learning / neuromuscular junction / recycling endosome / cognition / Golgi lumen / positive regulation of inflammatory response / positive regulation of interleukin-6 production / neuron cellular homeostasis / positive regulation of non-canonical NF-kappaB signal transduction / endocytosis / Regulation of Insulin-like Growth Factor (IGF) transport and uptake by Insulin-like Growth Factor Binding Proteins (IGFBPs) / cellular response to amyloid-beta / G2/M transition of mitotic cell cycle / positive regulation of tumor necrosis factor production / neuron projection development / cell-cell junction / synaptic vesicle / Platelet degranulation / apical part of cell Similarity search - Function | ||||||||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||||||||
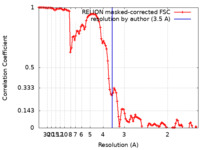
| Method | helical reconstruction / cryo EM / Resolution: 3.5 Å | ||||||||||||||||||
 Authors Authors | Zielinski M / Peralta Reyes FS / Gremer L / Schemmert S / Frieg B / Willuweit A / Donner L / Elvers M / Nilsson LNG / Syvanen S ...Zielinski M / Peralta Reyes FS / Gremer L / Schemmert S / Frieg B / Willuweit A / Donner L / Elvers M / Nilsson LNG / Syvanen S / Sehlin D / Ingelsson M / Willbold D / Schroeder GF | ||||||||||||||||||
| Funding support |  Germany, Germany,  Sweden, 5 items Sweden, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Nat Neurosci / Year: 2023 Journal: Nat Neurosci / Year: 2023Title: Cryo-EM of Aβ fibrils from mouse models find tg-APP fibrils resemble those found in patients with sporadic Alzheimer's disease. Authors: Mara Zielinski / Fernanda S Peralta Reyes / Lothar Gremer / Sarah Schemmert / Benedikt Frieg / Luisa U Schäfer / Antje Willuweit / Lili Donner / Margitta Elvers / Lars N G Nilsson / Stina ...Authors: Mara Zielinski / Fernanda S Peralta Reyes / Lothar Gremer / Sarah Schemmert / Benedikt Frieg / Luisa U Schäfer / Antje Willuweit / Lili Donner / Margitta Elvers / Lars N G Nilsson / Stina Syvänen / Dag Sehlin / Martin Ingelsson / Dieter Willbold / Gunnar F Schröder /     Abstract: The use of transgenic mice displaying amyloid-β (Aβ) brain pathology has been essential for the preclinical assessment of new treatment strategies for Alzheimer's disease. However, the properties ...The use of transgenic mice displaying amyloid-β (Aβ) brain pathology has been essential for the preclinical assessment of new treatment strategies for Alzheimer's disease. However, the properties of Aβ in such mice have not been systematically compared to Aβ in the brains of patients with Alzheimer's disease. Here, we determined the structures of nine ex vivo Aβ fibrils from six different mouse models by cryogenic-electron microscopy. We found novel Aβ fibril structures in the APP/PS1, ARTE10 and tg-SwDI models, whereas the human type II filament fold was found in the ARTE10, tg-APP and APP23 models. The tg-APP mice showed an Aβ fibril whose structure resembles the human type I filament found in patients with sporadic Alzheimer's disease. A detailed assessment of the Aβ fibril structure is key to the selection of adequate mouse models for the preclinical development of novel plaque-targeting therapeutics and positron emission tomography imaging tracers in Alzheimer's disease. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16960.map.gz emd_16960.map.gz | 92.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16960-v30.xml emd-16960-v30.xml emd-16960.xml emd-16960.xml | 15.4 KB 15.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16960_fsc.xml emd_16960_fsc.xml | 10.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_16960.png emd_16960.png | 53.6 KB | ||
| Filedesc metadata |  emd-16960.cif.gz emd-16960.cif.gz | 5.4 KB | ||
| Others |  emd_16960_half_map_1.map.gz emd_16960_half_map_1.map.gz emd_16960_half_map_2.map.gz emd_16960_half_map_2.map.gz | 79.3 MB 79.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16960 http://ftp.pdbj.org/pub/emdb/structures/EMD-16960 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16960 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16960 | HTTPS FTP |
-Validation report
| Summary document |  emd_16960_validation.pdf.gz emd_16960_validation.pdf.gz | 190.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_16960_full_validation.pdf.gz emd_16960_full_validation.pdf.gz | 189.9 KB | Display | |
| Data in XML |  emd_16960_validation.xml.gz emd_16960_validation.xml.gz | 500 B | Display | |
| Data in CIF |  emd_16960_validation.cif.gz emd_16960_validation.cif.gz | 373 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16960 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16960 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16960 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16960 | HTTPS FTP |
-Related structure data
| Related structure data |  8oloMC  8ol2C  8ol3C  8ol5C  8ol6C  8ol7C  8olgC  8olnC  8olqC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_16960.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16960.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Ex vivo Abeta40 fibril from ARTE10 mouse brain. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.808 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Ex vivo Abeta40 fibril from ARTE10 mouse brain. Half map 1.
| File | emd_16960_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Ex vivo Abeta40 fibril from ARTE10 mouse brain. Half map 1. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Ex vivo Abeta40 fibril from ARTE10 mouse brain. Half map 2.
| File | emd_16960_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Ex vivo Abeta40 fibril from ARTE10 mouse brain. Half map 2. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Amyloid fibril of amyloid-beta
| Entire | Name: Amyloid fibril of amyloid-beta |
|---|---|
| Components |
|
-Supramolecule #1: Amyloid fibril of amyloid-beta
| Supramolecule | Name: Amyloid fibril of amyloid-beta / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 20 kDa/nm |
-Macromolecule #1: Amyloid-beta protein 42
| Macromolecule | Name: Amyloid-beta protein 42 / type: protein_or_peptide / ID: 1 / Number of copies: 10 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 4.520087 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DAEFRHDSGY EVHHQKLVFF AEDVGSNKGA IIGLMVGGVV IA UniProtKB: Amyloid-beta precursor protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Number grids imaged: 1 / Number real images: 9521 / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)