[English] 日本語
 Yorodumi
Yorodumi- EMDB-16664: Human heparan sulfate N-deacetylase-N-sulfotransferase 1 in compl... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human heparan sulfate N-deacetylase-N-sulfotransferase 1 in complex with calcium, 3'-phosphoadenosine-5'-phosphosulfate and nanobody nAb13 (composite map and model). | |||||||||||||||||||||
 Map data Map data | composite map of NDST1 nAb13 complex, created by combining locally refined maps for the NDST1 N-terminal and deacetylase domains, and deacetylase and sulfotransferase domains | |||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||
 Keywords Keywords | Deacetylase / Sulfotransferase / Heparan Sulfate / Carbohydrate / Glycosaminoglycan / Nanobody | |||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology information[heparan sulfate]-glucosamine N-sulfotransferase / heparan sulfate N-sulfotransferase activity / heparan sulfate N-deacetylase activity / N-acetylglucosamine deacetylase activity / heparin proteoglycan biosynthetic process / embryonic neurocranium morphogenesis / embryonic viscerocranium morphogenesis / HS-GAG biosynthesis / glycosaminoglycan metabolic process / heparan sulfate proteoglycan biosynthetic process ...[heparan sulfate]-glucosamine N-sulfotransferase / heparan sulfate N-sulfotransferase activity / heparan sulfate N-deacetylase activity / N-acetylglucosamine deacetylase activity / heparin proteoglycan biosynthetic process / embryonic neurocranium morphogenesis / embryonic viscerocranium morphogenesis / HS-GAG biosynthesis / glycosaminoglycan metabolic process / heparan sulfate proteoglycan biosynthetic process / cardiac septum development / deacetylase activity / respiratory gaseous exchange by respiratory system / polysaccharide biosynthetic process / positive regulation of smoothened signaling pathway / coronary vasculature development / Hydrolases; Acting on carbon-nitrogen bonds, other than peptide bonds; In linear amides / aorta development / midbrain development / forebrain development / fibroblast growth factor receptor signaling pathway / trans-Golgi network membrane / positive regulation of MAPK cascade / cell population proliferation / inflammatory response / Golgi membrane / Golgi apparatus Similarity search - Function | |||||||||||||||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.15 Å | |||||||||||||||||||||
 Authors Authors | Mycroft-West CJ / Wu L | |||||||||||||||||||||
| Funding support |  United Kingdom, 6 items United Kingdom, 6 items
| |||||||||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Structural and mechanistic characterization of bifunctional heparan sulfate N-deacetylase-N-sulfotransferase 1. Authors: Courtney J Mycroft-West / Sahar Abdelkarim / Helen M E Duyvesteyn / Neha S Gandhi / Mark A Skidmore / Raymond J Owens / Liang Wu /    Abstract: Heparan sulfate (HS) polysaccharides are major constituents of the extracellular matrix, which are involved in myriad structural and signaling processes. Mature HS polysaccharides contain complex, ...Heparan sulfate (HS) polysaccharides are major constituents of the extracellular matrix, which are involved in myriad structural and signaling processes. Mature HS polysaccharides contain complex, non-templated patterns of sulfation and epimerization, which mediate interactions with diverse protein partners. Complex HS modifications form around initial clusters of glucosamine-N-sulfate (GlcNS) on nascent polysaccharide chains, but the mechanistic basis underpinning incorporation of GlcNS itself into HS remains unclear. Here, we determine cryo-electron microscopy structures of human N-deacetylase-N-sulfotransferase (NDST)1, the bifunctional enzyme primarily responsible for initial GlcNS modification of HS. Our structures reveal the architecture of both NDST1 deacetylase and sulfotransferase catalytic domains, alongside a non-catalytic N-terminal domain. The two catalytic domains of NDST1 adopt a distinct back-to-back topology that limits direct cooperativity. Binding analyses, aided by activity-modulating nanobodies, suggest that anchoring of the substrate at the sulfotransferase domain initiates the NDST1 catalytic cycle, providing a plausible mechanism for cooperativity despite spatial domain separation. Our data shed light on key determinants of NDST1 activity, and describe tools to probe NDST1 function in vitro and in vivo. | |||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16664.map.gz emd_16664.map.gz | 299.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16664-v30.xml emd-16664-v30.xml emd-16664.xml emd-16664.xml | 15.6 KB 15.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_16664.png emd_16664.png | 66.5 KB | ||
| Filedesc metadata |  emd-16664.cif.gz emd-16664.cif.gz | 6.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16664 http://ftp.pdbj.org/pub/emdb/structures/EMD-16664 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16664 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16664 | HTTPS FTP |
-Related structure data
| Related structure data |  8chsMC  8ccyC  8cd0C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_16664.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16664.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | composite map of NDST1 nAb13 complex, created by combining locally refined maps for the NDST1 N-terminal and deacetylase domains, and deacetylase and sulfotransferase domains | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.73 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Human heparan sulfate N-deacetylase-N-sulfotransferase complex wi...
| Entire | Name: Human heparan sulfate N-deacetylase-N-sulfotransferase complex with nAb13 |
|---|---|
| Components |
|
-Supramolecule #1: Human heparan sulfate N-deacetylase-N-sulfotransferase complex wi...
| Supramolecule | Name: Human heparan sulfate N-deacetylase-N-sulfotransferase complex with nAb13 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #2: Nanobody nAb13 - all CA rigid fit model derived from nanobody nAb7
| Supramolecule | Name: Nanobody nAb13 - all CA rigid fit model derived from nanobody nAb7 type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: N-deacetylase-N-sulfotransferase 1
| Supramolecule | Name: N-deacetylase-N-sulfotransferase 1 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Nanobody nAb13 - all CA rigid fit model derived from nanobody nAb7
| Macromolecule | Name: Nanobody nAb13 - all CA rigid fit model derived from nanobody nAb7 type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 17.656316 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: QVQLVESGGG SVQAGGSLRL SCAASGFNVD DYAIGWFRQS PGKEREGVSC IGGDGTTYYE NSVKGRFTVS SDKRDNTVYL QMNNLRPED TAIYFCAADR SKYCVGKYFS TPSQYDFWGR GTHVTVSSEP KTPKPQPAAG PGGQHHHHHH GAEQKLISEE D LS |
-Macromolecule #2: Bifunctional heparan sulfate N-deacetylase/N-sulfotransferase 1
| Macromolecule | Name: Bifunctional heparan sulfate N-deacetylase/N-sulfotransferase 1 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO EC number: Hydrolases; Acting on carbon-nitrogen bonds, other than peptide bonds; In linear amides |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 92.464508 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: GSRTDPLVLV FVESLYSQLG QEVVAILESS RFKYRTEIAP GKGDMPTLTD KGRGRFALII YENILKYVNL DAWNRELLDK YCVAYGVGI IGFFKANENS LLSAQLKGFP LFLHSNLGLK DCSINPKSPL LYVTRPSEVE KGVLPGEDWT VFQSNHSTYE P VLLAKTRS ...String: GSRTDPLVLV FVESLYSQLG QEVVAILESS RFKYRTEIAP GKGDMPTLTD KGRGRFALII YENILKYVNL DAWNRELLDK YCVAYGVGI IGFFKANENS LLSAQLKGFP LFLHSNLGLK DCSINPKSPL LYVTRPSEVE KGVLPGEDWT VFQSNHSTYE P VLLAKTRS SESIPHLGAD AGLHAALHAT VVQDLGLHDG IQRVLFGNNL NFWLHKLVFV DAVAFLTGKR LSLPLDRYIL VD IDDIFVG KEGTRMKVED VKALFDTQNE LRAHIPNFTF NLGYSGKFFH TGTNAEDAGD DLLLSYVKEF WWFPHMWSHM QPH LFHNQS VLAEQMALNK KFAVEHGIPT DMGYAVAPHH SGVYPVHVQL YEAWKQVWSI RVTSTEEYPH LKPARYRRGF IHNG IMVLP RQTCGLFTHT IFYNEYPGGS SELDKIINGG ELFLTVLLNP ISIFMTHLSN YGNDRLGLYT FKHLVRFLHS WTNLR LQTL PPVQLAQKYF QIFSEEKDPL WQDPCEDKRH KDIWSKEKTC DRFPKLLIIG PQKTGTTALY LFLGMHPDLS SNYPSS ETF EEIQFFNGHN YHKGIDWYME FFPIPSNTTS DFYFEKSANY FDSEVAPRRA AALLPKAKVL TILINPADRA YSWYQHQ RA HDDPVALKYT FHEVITAGSD ASSKLRALQN RCLVPGWYAT HIERWLSAYH ANQILVLDGK LLRTEPAKVM DMVQKFLG V TNTIDYHKTL AFDPKKGFWC QLLEGGKTKC LGKSKGRKYP EMDLDSRAFL KDYYRDHNIE LSKLLYKMGQ TLPTWLRED LQNTR UniProtKB: Bifunctional heparan sulfate N-deacetylase/N-sulfotransferase 1 |
-Macromolecule #3: ADENOSINE-3'-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-3'-5'-DIPHOSPHATE / type: ligand / ID: 3 / Number of copies: 1 / Formula: A3P |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information | 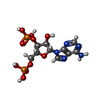 ChemComp-A3P: |
-Macromolecule #4: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 4 / Number of copies: 1 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Macromolecule #5: water
| Macromolecule | Name: water / type: ligand / ID: 5 / Number of copies: 1 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.4 mg/mL |
|---|---|
| Buffer | pH: 6.5 |
| Vitrification | Cryogen name: ETHANE |
| Details | Monodisperse sample from size exclusion |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.6 µm / Nominal defocus min: 1.2 µm / Nominal magnification: 165000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: INSILICO MODEL |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.15 Å / Resolution method: FSC 0.143 CUT-OFF Details: composite map made by combining locally refined volumes of NDST1 N-terminal+deacetylase, and NDST1 deacetylase+sulfotransferase domains. Resolution reported for composite map is the ...Details: composite map made by combining locally refined volumes of NDST1 N-terminal+deacetylase, and NDST1 deacetylase+sulfotransferase domains. Resolution reported for composite map is the resolution of the lowest locally refined component. Number images used: 87798 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)