+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | HK97 Portal Protein In situ (prohead II) | |||||||||
 Map data Map data | HK97 bacteriophage portal protein in prohead II as part of packaging complex. Symmetry = C12. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | portal / HK97 / bacteriophage / packaging / VIRAL PROTEIN | |||||||||
| Function / homology | Phage portal protein, HK97 / Bacteriophage/Gene transfer agent portal protein / Phage portal protein / viral capsid / Portal protein Function and homology information Function and homology information | |||||||||
| Biological species |  Hendrixvirus Hendrixvirus | |||||||||
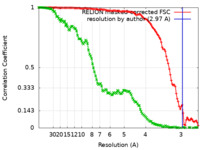
| Method | single particle reconstruction / cryo EM / Resolution: 2.97 Å | |||||||||
 Authors Authors | Hawkins DEDP / Antson AA | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2023 Journal: Nucleic Acids Res / Year: 2023Title: Insights into a viral motor: the structure of the HK97 packaging termination assembly. Authors: Dorothy E D P Hawkins / Oliver W Bayfield / Herman K H Fung / Daniel N Grba / Alexis Huet / James F Conway / Alfred A Antson /    Abstract: Double-stranded DNA viruses utilise machinery, made of terminase proteins, to package viral DNA into the capsid. For cos bacteriophage, a defined signal, recognised by small terminase, flanks each ...Double-stranded DNA viruses utilise machinery, made of terminase proteins, to package viral DNA into the capsid. For cos bacteriophage, a defined signal, recognised by small terminase, flanks each genome unit. Here we present the first structural data for a cos virus DNA packaging motor, assembled from the bacteriophage HK97 terminase proteins, procapsids encompassing the portal protein, and DNA containing a cos site. The cryo-EM structure is consistent with the packaging termination state adopted after DNA cleavage, with DNA density within the large terminase assembly ending abruptly at the portal protein entrance. Retention of the large terminase complex after cleavage of the short DNA substrate suggests that motor dissociation from the capsid requires headful pressure, in common with pac viruses. Interestingly, the clip domain of the 12-subunit portal protein does not adhere to C12 symmetry, indicating asymmetry induced by binding of the large terminase/DNA. The motor assembly is also highly asymmetric, showing a ring of 5 large terminase monomers, tilted against the portal. Variable degrees of extension between N- and C-terminal domains of individual subunits suggest a mechanism of DNA translocation driven by inter-domain contraction and relaxation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16614.map.gz emd_16614.map.gz | 116.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16614-v30.xml emd-16614-v30.xml emd-16614.xml emd-16614.xml | 16.1 KB 16.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16614_fsc.xml emd_16614_fsc.xml emd_16614_fsc_2.xml emd_16614_fsc_2.xml | 11.3 KB 14.6 KB | Display Display |  FSC data file FSC data file |
| Images |  emd_16614.png emd_16614.png | 68.6 KB | ||
| Masks |  emd_16614_msk_1.map emd_16614_msk_1.map | 125 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-16614.cif.gz emd-16614.cif.gz | 6.1 KB | ||
| Others |  emd_16614_half_map_1.map.gz emd_16614_half_map_1.map.gz emd_16614_half_map_2.map.gz emd_16614_half_map_2.map.gz | 98.1 MB 98.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16614 http://ftp.pdbj.org/pub/emdb/structures/EMD-16614 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16614 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16614 | HTTPS FTP |
-Validation report
| Summary document |  emd_16614_validation.pdf.gz emd_16614_validation.pdf.gz | 1.1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_16614_full_validation.pdf.gz emd_16614_full_validation.pdf.gz | 1.1 MB | Display | |
| Data in XML |  emd_16614_validation.xml.gz emd_16614_validation.xml.gz | 18.6 KB | Display | |
| Data in CIF |  emd_16614_validation.cif.gz emd_16614_validation.cif.gz | 24.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16614 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16614 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16614 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16614 | HTTPS FTP |
-Related structure data
| Related structure data |  8cezMC  8cfaC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_16614.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16614.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | HK97 bacteriophage portal protein in prohead II as part of packaging complex. Symmetry = C12. | ||||||||||||||||||||||||||||||||||||
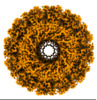
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.34 Å | ||||||||||||||||||||||||||||||||||||
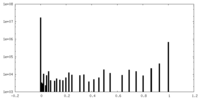
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_16614_msk_1.map emd_16614_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
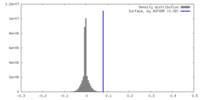
| Density Histograms |
-Half map: HK97 bacteriophage portal protein in prohead II as...
| File | emd_16614_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | HK97 bacteriophage portal protein in prohead II as part of packaging complex. Symmetry = C12. | ||||||||||||
| Projections & Slices |
| ||||||||||||
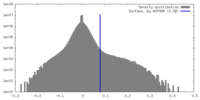
| Density Histograms |
-Half map: HK97 bacteriophage portal protein in prohead II as...
| File | emd_16614_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | HK97 bacteriophage portal protein in prohead II as part of packaging complex. Symmetry = C12. | ||||||||||||
| Projections & Slices |
| ||||||||||||
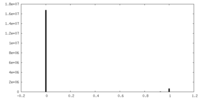
| Density Histograms |
- Sample components
Sample components
-Entire : Hendrixvirus
| Entire | Name:  Hendrixvirus Hendrixvirus |
|---|---|
| Components |
|
-Supramolecule #1: Hendrixvirus
| Supramolecule | Name: Hendrixvirus / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all Details: Proheads were produced by infection of E. coli 594 cells with HK97 amber mutant amC2, propagated using Escherichia LE392 cells. NCBI-ID: 2169654 / Sci species name: Hendrixvirus / Sci species strain: HK97 / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Escherichia phage EcSzw-2 (virus) / Strain: 594 Escherichia phage EcSzw-2 (virus) / Strain: 594 |
| Molecular weight | Theoretical: 560 KDa |
-Macromolecule #1: Portal protein
| Macromolecule | Name: Portal protein / type: protein_or_peptide / ID: 1 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Hendrixvirus Hendrixvirus |
| Molecular weight | Theoretical: 47.318488 KDa |
| Recombinant expression | Organism:  Escherichia phage EcSzw-2 (virus) Escherichia phage EcSzw-2 (virus) |
| Sequence | String: MEEPKYTIDL RTNNGWWARL KSWFVGGRLV TPNQGSQTGP VSAHGYLGDS SINDERILQI STVWRCVSLI STLTACLPLD VFETDQNDN RKKVDLSNPL ARLLRYSPNQ YMTAQEFREA MTMQLCFYGN AYALVDRNSA GDVISLLPLQ SANMDVKLVG K KVVYRYQR ...String: MEEPKYTIDL RTNNGWWARL KSWFVGGRLV TPNQGSQTGP VSAHGYLGDS SINDERILQI STVWRCVSLI STLTACLPLD VFETDQNDN RKKVDLSNPL ARLLRYSPNQ YMTAQEFREA MTMQLCFYGN AYALVDRNSA GDVISLLPLQ SANMDVKLVG K KVVYRYQR DSEYADFSQK EIFHLKGFGF TGLVGLSPIA FACKSAGVAV AMEDQQRDFF ANGAKSPQIL STGEKVLTEQ QR SQVEENF KEIAGGPVKK RLWILEAGFS TSAIGVTPQD AEMMASRKFQ VSELARFFGV PPHLVGDVEK STSWGSGIEQ QNL GFLQYT LQPYISRWEN SIQRWLIPSK DVGRLHAEHN LDGLLRGDSA SRAAFMKAMG ESGLRTINEM RRTDNMPPLP GGDV AMRQA QYVPITDLGT NKEPRNNGA UniProtKB: Portal protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Details: final buffer = 20 mM Tris pH 7.5, 10 mM MgCl2, 50 mM KGlu, 5 mM ATP-gamma-S |
|---|---|
| Grid | Model: Quantifoil / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 0.2 / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 60 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Instrument: FEI VITROBOT MARK IV |
| Details | Packaging reactions 30 nM hk97 proheads, 2.5 micromolar large terminase, 5 micromolar small terminase and 30 nM DNA, 75 micromolar ATP were incubated for 2 mins then 5 micromolar ATP-gamma-S was added. |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)