[English] 日本語
 Yorodumi
Yorodumi- EMDB-16545: Structure of human mitochondrial CCA-adding enzyme in complex wit... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
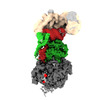
| Title | Structure of human mitochondrial CCA-adding enzyme in complex with mitochondrial pre-tRNA-Ile | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | RNA maturation / RNA modification / mitochondrial tRNA / RNA methyltransferase / MRPP1 / MRPP2 / MRPP3 / RNA binding protein / m6A / RNA binding / RNA recognition | |||||||||
| Function / homology |  Function and homology information Function and homology information5'-3' RNA polymerase activity / cardiolipin dehydrogenase (NAD+) activity / brexanolone metabolic process / isoursodeoxycholate 7-beta-dehydrogenase (NAD+) activity / ursodeoxycholate 7-beta-dehydrogenase (NAD+) activity / 3-hydroxy-2-methylbutyryl-CoA dehydrogenase / 3-hydroxy-2-methylbutyryl-CoA dehydrogenase activity / mitochondrial tRNA methylation / tRNA (adenine9-N1)-methyltransferase / mitochondrial tRNA processing ...5'-3' RNA polymerase activity / cardiolipin dehydrogenase (NAD+) activity / brexanolone metabolic process / isoursodeoxycholate 7-beta-dehydrogenase (NAD+) activity / ursodeoxycholate 7-beta-dehydrogenase (NAD+) activity / 3-hydroxy-2-methylbutyryl-CoA dehydrogenase / 3-hydroxy-2-methylbutyryl-CoA dehydrogenase activity / mitochondrial tRNA methylation / tRNA (adenine9-N1)-methyltransferase / mitochondrial tRNA processing / tRNA (adenine(9)-N1)-methyltransferase activity / mitochondrial ribonuclease P complex / mitochondrial tRNA 5'-end processing / chenodeoxycholate 7-alpha-dehydrogenase (NAD+) activity / rRNA processing in the mitochondrion / tRNA processing in the mitochondrion / tRNA modification in the mitochondrion / tRNA (guanine9-N1)-methyltransferase / tRNA (guanosine(9)-N1)-methyltransferase activity / 7alpha-hydroxysteroid dehydrogenase / 17-beta-hydroxysteroid dehydrogenase (NAD+) activity / mitochondrial tRNA 3'-end processing / cholate 7-alpha-dehydrogenase (NAD+) activity / mitochondrial RNA 5'-end processing / C21-steroid hormone metabolic process / CCA tRNA nucleotidyltransferase / tRNA surveillance / CCA tRNA nucleotidyltransferase activity / CCACCA tRNA nucleotidyltransferase activity / tRNA 3'-terminal CCA addition / tRNA methyltransferase complex / 3-hydroxyacyl-CoA dehydrogenase / tRNA 3'-end processing / L-isoleucine catabolic process / 3alpha(17beta)-hydroxysteroid dehydrogenase (NAD+) / 3alpha(or 20beta)-hydroxysteroid dehydrogenase / (3S)-3-hydroxyacyl-CoA dehydrogenase (NAD+) activity / androstan-3-alpha,17-beta-diol dehydrogenase (NAD+) activity / testosterone dehydrogenase (NAD+) activity / positive regulation of mitochondrial translation / bile acid biosynthetic process / 17beta-estradiol 17-dehydrogenase / estradiol 17-beta-dehydrogenase [NAD(P)+] activity / Branched-chain amino acid catabolism / estrogen metabolic process / tRNA processing in the nucleus / fatty acid beta-oxidation / androgen metabolic process / mitochondrial nucleoid / Mitochondrial protein degradation / rescue of stalled cytosolic ribosome / Transferases; Transferring one-carbon groups; Methyltransferases / lipid metabolic process / mitochondrion organization / fatty acid metabolic process / mRNA processing / protein homotetramerization / tRNA binding / mitochondrial matrix / protein homodimerization activity / mitochondrion / RNA binding / nucleoplasm / ATP binding / metal ion binding / identical protein binding / nucleus / plasma membrane / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.14 Å | |||||||||
 Authors Authors | MEYNIER V / HARDWICK S / CATALA M / ROSKE J / OERUM S / CHIRGADZE D / BARRAUD P / LUISI B / TISNE C | |||||||||
| Funding support |  France, 1 items France, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Structural basis for human mitochondrial tRNA maturation. Authors: Vincent Meynier / Steven W Hardwick / Marjorie Catala / Johann J Roske / Stephanie Oerum / Dimitri Y Chirgadze / Pierre Barraud / Wyatt W Yue / Ben F Luisi / Carine Tisné /   Abstract: The human mitochondrial genome is transcribed into two RNAs, containing mRNAs, rRNAs and tRNAs, all dedicated to produce essential proteins of the respiratory chain. The precise excision of tRNAs by ...The human mitochondrial genome is transcribed into two RNAs, containing mRNAs, rRNAs and tRNAs, all dedicated to produce essential proteins of the respiratory chain. The precise excision of tRNAs by the mitochondrial endoribonucleases (mt-RNase), P and Z, releases all RNA species from the two RNA transcripts. The tRNAs then undergo 3'-CCA addition. In metazoan mitochondria, RNase P is a multi-enzyme assembly that comprises the endoribonuclease PRORP and a tRNA methyltransferase subcomplex. The requirement for this tRNA methyltransferase subcomplex for mt-RNase P cleavage activity, as well as the mechanisms of pre-tRNA 3'-cleavage and 3'-CCA addition, are still poorly understood. Here, we report cryo-EM structures that visualise four steps of mitochondrial tRNA maturation: 5' and 3' tRNA-end processing, methylation and 3'-CCA addition, and explain the defined sequential order of the tRNA processing steps. The methyltransferase subcomplex recognises the pre-tRNA in a distinct mode that can support tRNA-end processing and 3'-CCA addition, likely resulting from an evolutionary adaptation of mitochondrial tRNA maturation complexes to the structurally-fragile mitochondrial tRNAs. This subcomplex can also ensure a tRNA-folding quality-control checkpoint before the sequential docking of the maturation enzymes. Altogether, our study provides detailed molecular insight into RNA-transcript processing and tRNA maturation in human mitochondria. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16545.map.gz emd_16545.map.gz | 206.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16545-v30.xml emd-16545-v30.xml emd-16545.xml emd-16545.xml | 22.3 KB 22.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_16545.png emd_16545.png | 62.9 KB | ||
| Filedesc metadata |  emd-16545.cif.gz emd-16545.cif.gz | 7.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16545 http://ftp.pdbj.org/pub/emdb/structures/EMD-16545 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16545 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16545 | HTTPS FTP |
-Related structure data
| Related structure data |  8cbmMC  8cbkC  8cblC  8cboC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_16545.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16545.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.652 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Human mitochondrial CCA-adding enzyme in complex with mitochondri...
| Entire | Name: Human mitochondrial CCA-adding enzyme in complex with mitochondrial pre-tRNAIle |
|---|---|
| Components |
|
-Supramolecule #1: Human mitochondrial CCA-adding enzyme in complex with mitochondri...
| Supramolecule | Name: Human mitochondrial CCA-adding enzyme in complex with mitochondrial pre-tRNAIle type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 270 KDa |
-Macromolecule #1: 3-hydroxyacyl-CoA dehydrogenase type-2
| Macromolecule | Name: 3-hydroxyacyl-CoA dehydrogenase type-2 / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO / EC number: 3-hydroxyacyl-CoA dehydrogenase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 26.947021 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAAACRSVKG LVAVITGGAS GLGLATAERL VGQGASAVLL DLPNSGGEAQ AKKLGNNCVF APADVTSEKD VQTALALAKG KFGRVDVAV NCAGIAVASK TYNLKKGQTH TLEDFQRVLD VNLMGTFNVI RLVAGEMGQN EPDQGGQRGV IINTASVAAF E GQVGQAAY ...String: MAAACRSVKG LVAVITGGAS GLGLATAERL VGQGASAVLL DLPNSGGEAQ AKKLGNNCVF APADVTSEKD VQTALALAKG KFGRVDVAV NCAGIAVASK TYNLKKGQTH TLEDFQRVLD VNLMGTFNVI RLVAGEMGQN EPDQGGQRGV IINTASVAAF E GQVGQAAY SASKGGIVGM TLPIARDLAP IGIRVMTIAP GLFGTPLLTS LPEKVCNFLA SQVPFPSRLG DPAEYAHLVQ AI IENPFLN GEVIRLDGAI RMQP UniProtKB: 3-hydroxyacyl-CoA dehydrogenase type-2 |
-Macromolecule #2: CCA tRNA nucleotidyltransferase 1, mitochondrial
| Macromolecule | Name: CCA tRNA nucleotidyltransferase 1, mitochondrial / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO / EC number: CCA tRNA nucleotidyltransferase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 44.989277 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: TEGLKSLTEL FVKENHELRI AGGAVRDLLN GVKPQDIDFA TTATPTQMKE MFQSAGIRMI NNRGEKHGTI TARLHEENFE ITTLRIDVT TDGRHAEVEF TTDWQKDAER RDLTINSMFL GFDGTLFDYF NGYEDLKNKK VRFVGHAKQR IQEDYLRILR Y FRFYGRIV ...String: TEGLKSLTEL FVKENHELRI AGGAVRDLLN GVKPQDIDFA TTATPTQMKE MFQSAGIRMI NNRGEKHGTI TARLHEENFE ITTLRIDVT TDGRHAEVEF TTDWQKDAER RDLTINSMFL GFDGTLFDYF NGYEDLKNKK VRFVGHAKQR IQEDYLRILR Y FRFYGRIV DKPGDHDPET LEAIAENAKG LAGISGERIW VELKKILVGN HVNHLIHLIY DLDVAPYIGL PANASLEEFD KV SKNVDGF SPKPVTLLAS LFKVQDDVTK LDLRLKIAKE EKNLGLFIVK NRKDLIKATD SSDPLKPYQD FIIDSREPDA TTR VCELLK YQGEHCLLKE MQQWSIPPFP VSGHDIRKVG ISSGKEIGAL LQQLREQWKK SGYQMEKDEL LSYIKKT UniProtKB: CCA tRNA nucleotidyltransferase 1, mitochondrial |
-Macromolecule #3: tRNA methyltransferase 10 homolog C
| Macromolecule | Name: tRNA methyltransferase 10 homolog C / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO EC number: Transferases; Transferring one-carbon groups; Methyltransferases |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 48.007793 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MHHHHHHSSG VDLGTENLYF QSMSSKIPAV TYPKNESTPP SEELELDKWK TTMKSSVQEE CVSTISSSKD EDPLAATREF IEMWRLLGR EVPEHITEEE LKTLMECVSN TAKKKYLKYL YTKEKVKKAR QIKKEMKAAA REEAKNIKLL ETTEEDKQKN F LFLRLWDR ...String: MHHHHHHSSG VDLGTENLYF QSMSSKIPAV TYPKNESTPP SEELELDKWK TTMKSSVQEE CVSTISSSKD EDPLAATREF IEMWRLLGR EVPEHITEEE LKTLMECVSN TAKKKYLKYL YTKEKVKKAR QIKKEMKAAA REEAKNIKLL ETTEEDKQKN F LFLRLWDR NMDIAMGWKG AQAMQFGQPL VFDMAYENYM KRKELQNTVS QLLESEGWNR RNVDPFHIYF CNLKIDGALH RE LVKRYQE KWDKLLLTST EKSHVDLFPK DSIIYLTADS PNVMTTFRHD KVYVIGSFVD KSMQPGTSLA KAKRLNLATE CLP LDKYLQ WEIGNKNLTL DQMIRILLCL KNNGNWQEAL QFVPKRKHTG FLEISQHSQE FINRLKKAKT AENLYFQSHH HHHH DYKDD DDK UniProtKB: tRNA methyltransferase 10 homolog C |
-Macromolecule #4: Mitochondrial Precursor tRNA-Ile(0,0)
| Macromolecule | Name: Mitochondrial Precursor tRNA-Ile(0,0) / type: rna / ID: 4 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 22.138164 KDa |
| Sequence | String: GGAAAUAUGU CUGAUAAAAG AGUUACUUUG AUAGAGUAAA UAAUAGGAGC UUAAACCCCC UUAUUUCCA |
-Macromolecule #5: NICOTINAMIDE-ADENINE-DINUCLEOTIDE
| Macromolecule | Name: NICOTINAMIDE-ADENINE-DINUCLEOTIDE / type: ligand / ID: 5 / Number of copies: 4 / Formula: NAD |
|---|---|
| Molecular weight | Theoretical: 663.425 Da |
| Chemical component information |  ChemComp-NAD: |
-Macromolecule #6: CYTIDINE-5'-DIPHOSPHATE
| Macromolecule | Name: CYTIDINE-5'-DIPHOSPHATE / type: ligand / ID: 6 / Number of copies: 1 / Formula: CDP |
|---|---|
| Molecular weight | Theoretical: 403.176 Da |
| Chemical component information | 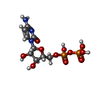 ChemComp-CDP: |
-Macromolecule #7: S-ADENOSYL-L-HOMOCYSTEINE
| Macromolecule | Name: S-ADENOSYL-L-HOMOCYSTEINE / type: ligand / ID: 7 / Number of copies: 1 / Formula: SAH |
|---|---|
| Molecular weight | Theoretical: 384.411 Da |
| Chemical component information |  ChemComp-SAH: |
-Macromolecule #8: water
| Macromolecule | Name: water / type: ligand / ID: 8 / Number of copies: 8 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 Component:
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 3 sec. | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 42.2 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: DIFFRACTION / Nominal defocus max: 2.2 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 130000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: AB INITIO MODEL / Overall B value: 60.7 / Target criteria: Cross-correlation coefficient |
|---|---|
| Output model |  PDB-8cbm: |
 Movie
Movie Controller
Controller





















 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)





















