[English] 日本語
 Yorodumi
Yorodumi- EMDB-16531: Cryo-EM structure of the Cora homohexamer from Galleria mellonell... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the Cora homohexamer from Galleria mellonella saliva | ||||||||||||||||||||||||
 Map data Map data | |||||||||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||||||||
 Keywords Keywords | Galleria mellonella / Wax worm saliva / Plastic degradation / Polyethylene degradation / PEases / Hexamerin / Hemocyanin/phenoloxidase superfamily / Metal binding / UNKNOWN FUNCTION | ||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationArthropod hemocyanins / insect LSPs signature 1. / Hemocyanin, N-terminal / Hemocyanin, N-terminal domain superfamily / Hemocyanin, all-alpha domain / Hemocyanin, C-terminal domain superfamily / Hemocyanin/hexamerin middle domain / Hemocyanin, C-terminal / Hemocyanin, copper containing domain / Hemocyanin, ig-like domain / Hemocyanin/hexamerin ...Arthropod hemocyanins / insect LSPs signature 1. / Hemocyanin, N-terminal / Hemocyanin, N-terminal domain superfamily / Hemocyanin, all-alpha domain / Hemocyanin, C-terminal domain superfamily / Hemocyanin/hexamerin middle domain / Hemocyanin, C-terminal / Hemocyanin, copper containing domain / Hemocyanin, ig-like domain / Hemocyanin/hexamerin / Di-copper centre-containing domain superfamily / Immunoglobulin E-set Similarity search - Domain/homology | ||||||||||||||||||||||||
| Biological species |  Galleria mellonella (greater wax moth) Galleria mellonella (greater wax moth) | ||||||||||||||||||||||||
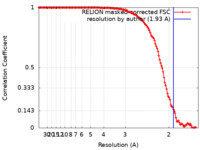
| Method | single particle reconstruction / cryo EM / Resolution: 1.93 Å | ||||||||||||||||||||||||
 Authors Authors | Spinola-Amilibia M / Arias-Palomo E / Araujo-Bazan L / Berger JM | ||||||||||||||||||||||||
| Funding support |  Germany, Germany,  Spain, Spain,  Belgium, 7 items Belgium, 7 items
| ||||||||||||||||||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2023 Journal: Sci Adv / Year: 2023Title: Plastic degradation by insect hexamerins: Near-atomic resolution structures of the polyethylene-degrading proteins from the wax worm saliva. Authors: Mercedes Spínola-Amilibia / Ramiro Illanes-Vicioso / Elena Ruiz-López / Pere Colomer-Vidal / Francisco Rodriguez-Ventura / Rosa Peces Pérez / Clemente F Arias / Tomas Torroba / Maria ...Authors: Mercedes Spínola-Amilibia / Ramiro Illanes-Vicioso / Elena Ruiz-López / Pere Colomer-Vidal / Francisco Rodriguez-Ventura / Rosa Peces Pérez / Clemente F Arias / Tomas Torroba / Maria Solà / Ernesto Arias-Palomo / Federica Bertocchini /  Abstract: Plastic waste management is a pressing ecological, social, and economic challenge. The saliva of the lepidopteran larvae is capable of oxidizing and depolymerizing polyethylene in hours at room ...Plastic waste management is a pressing ecological, social, and economic challenge. The saliva of the lepidopteran larvae is capable of oxidizing and depolymerizing polyethylene in hours at room temperature. Here, we analyze by cryo-electron microscopy (cryo-EM) 's saliva directly from the native source. The three-dimensional reconstructions reveal that the buccal secretion is mainly composed of four hexamerins belonging to the hemocyanin/phenoloxidase family, renamed Demetra, Cibeles, Ceres, and a previously unidentified factor termed Cora. Functional assays show that this factor, as its counterparts Demetra and Ceres, is also able to oxidize and degrade polyethylene. The cryo-EM data and the x-ray analysis from purified fractions show that they self-assemble primarily into three macromolecular complexes with striking structural differences that likely modulate their activity. Overall, these results establish the ground to further explore the hexamerins' functionalities, their role in vivo, and their eventual biotechnological application. | ||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16531.map.gz emd_16531.map.gz | 186.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16531-v30.xml emd-16531-v30.xml emd-16531.xml emd-16531.xml | 24 KB 24 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16531_fsc.xml emd_16531_fsc.xml | 13.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_16531.png emd_16531.png | 190.1 KB | ||
| Masks |  emd_16531_msk_1.map emd_16531_msk_1.map | 199.6 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-16531.cif.gz emd-16531.cif.gz | 6.9 KB | ||
| Others |  emd_16531_additional_1.map.gz emd_16531_additional_1.map.gz emd_16531_half_map_1.map.gz emd_16531_half_map_1.map.gz emd_16531_half_map_2.map.gz emd_16531_half_map_2.map.gz | 157.7 MB 158.2 MB 158.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16531 http://ftp.pdbj.org/pub/emdb/structures/EMD-16531 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16531 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16531 | HTTPS FTP |
-Related structure data
| Related structure data |  8canMC  8ca9C  8cadC  8po9C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_16531.map.gz / Format: CCP4 / Size: 199.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16531.map.gz / Format: CCP4 / Size: 199.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.81595 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_16531_msk_1.map emd_16531_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #1
| File | emd_16531_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_16531_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_16531_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cora homocomplex
| Entire | Name: Cora homocomplex |
|---|---|
| Components |
|
-Supramolecule #1: Cora homocomplex
| Supramolecule | Name: Cora homocomplex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 / Details: Homohexamer |
|---|---|
| Source (natural) | Organism:  Galleria mellonella (greater wax moth) Galleria mellonella (greater wax moth) |
-Macromolecule #1: basic juvenile hormone-suppressible protein 1
| Macromolecule | Name: basic juvenile hormone-suppressible protein 1 / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Galleria mellonella (greater wax moth) Galleria mellonella (greater wax moth) |
| Molecular weight | Theoretical: 88.394211 KDa |
| Sequence | String: MVVTMRLVVA AVLLAAVAAS VVHDELKNIV ITKEPMKNMD MKSKEMCILK LMNHILQPTM YEDVREVAKA WVLEENEDKY MKMEAVKEF INTYKMGMLP RGEVFVHMDH KHVEEAVKVF KLLYFANDFD VFLKTACWLR ERINGGMFVY ALTAAIFHRS D CSGIKIPA ...String: MVVTMRLVVA AVLLAAVAAS VVHDELKNIV ITKEPMKNMD MKSKEMCILK LMNHILQPTM YEDVREVAKA WVLEENEDKY MKMEAVKEF INTYKMGMLP RGEVFVHMDH KHVEEAVKVF KLLYFANDFD VFLKTACWLR ERINGGMFVY ALTAAIFHRS D CSGIKIPA PYEIYPYLFV DSNILHKAFM MKMSKAAMDP VMKNYYGIKV KDNSMVIIDW RKGLRHTMSE FDRTSYFTED ID LNTYLYY MHMSYPYWMN EDMYRVNKER RGEAMWYGYQ QLQARLRLER LSHHMCDLKP LDLDGTLDEG YWPKILLHTG DEM PVRYNK MKLTNENNIK YRLLLEDNKR LIRDGIKKGH MAMHDGTTVS LKKPDDIENL CRIVLGGFVS KDDHKGKSSI WRNL AKTML SYGTYNMGKY TYIPTAADMY STALRDPGMW KMLKLISEYF IMFKEMLPKY TREELDFPGV KIEQVTTDKL VTFMD EYDV DITNAVYLDH DEMQKHRSDM MYVARMHRLN HQPFKITIDV ASDKAVECVV RVFLGPKLDC MGRFTSVNDK RNDMVE IDS FLYKLETGKN TIVRDSLEMN NVIKERPWSR NNWAMDPSGG QKAQDNWWYK SRIGFPHRLL LPMGSHGGMP YQMFVIV TP VRAGMSLPSI DMNTAKERKA CRWTVCMDTM PLGFPFDRPI DETNFYTKNM KFHDVMVYTK DLAMSNMVKD VDMSEMVM K RDDLTYLDKD MLVKRSYKSV MMMSGDDMTH M UniProtKB: Basic juvenile hormone-suppressible protein 1 |
-Macromolecule #2: TRYPTOPHAN
| Macromolecule | Name: TRYPTOPHAN / type: ligand / ID: 2 / Number of copies: 6 / Formula: TRP |
|---|---|
| Molecular weight | Theoretical: 204.225 Da |
| Chemical component information |  ChemComp-TRP: |
-Macromolecule #3: COPPER (II) ION
| Macromolecule | Name: COPPER (II) ION / type: ligand / ID: 3 / Number of copies: 24 / Formula: CU |
|---|---|
| Molecular weight | Theoretical: 63.546 Da |
| Chemical component information |  ChemComp-CU: |
-Macromolecule #4: SODIUM ION
| Macromolecule | Name: SODIUM ION / type: ligand / ID: 4 / Number of copies: 3 |
|---|---|
| Molecular weight | Theoretical: 22.99 Da |
-Macromolecule #5: water
| Macromolecule | Name: water / type: ligand / ID: 5 / Number of copies: 3170 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Component:
| |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 2 sec. / Pretreatment - Atmosphere: AIR / Details: 25 mA | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 298 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number real images: 6386 / Average exposure time: 5.5 sec. / Average electron dose: 58.3 e/Å2 Details: Images were collected in super-resolution mode (pixel size of 0.543 angstrom) |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.6 µm / Nominal defocus min: 1.1 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL / Overall B value: 39.074 |
|---|---|
| Output model |  PDB-8can: |
 Movie
Movie Controller
Controller





 Z
Z Y
Y X
X