[English] 日本語
 Yorodumi
Yorodumi- EMDB-16264: Cryo-EM structure of the Hfq-Crc-amiE translation repression assembly. -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the Hfq-Crc-amiE translation repression assembly. | |||||||||
 Map data Map data | Locally filtered cryo-EM map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | co-transcriptional RNA folding / Crc / metabolic regulation / ribonucleoprotein assembly / RNA chaperone Hfq / translational regulation / RNA BINDING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationdouble-stranded DNA 3'-5' DNA exonuclease activity / exodeoxyribonuclease III / DNA repair / regulation of DNA-templated transcription / RNA binding / metal ion binding Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||
 Authors Authors | Dendooven T / Luisi BF | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: EMBO J / Year: 2023 Journal: EMBO J / Year: 2023Title: Translational regulation by Hfq-Crc assemblies emerges from polymorphic ribonucleoprotein folding. Authors: Tom Dendooven / Elisabeth Sonnleitner / Udo Bläsi / Ben F Luisi /   Abstract: The widely occurring bacterial RNA chaperone Hfq is a key factor in the post-transcriptional control of hundreds of genes in Pseudomonas aeruginosa. How this broadly acting protein can contribute to ...The widely occurring bacterial RNA chaperone Hfq is a key factor in the post-transcriptional control of hundreds of genes in Pseudomonas aeruginosa. How this broadly acting protein can contribute to the regulatory requirements of many different genes remains puzzling. Here, we describe cryo-EM structures of higher order assemblies formed by Hfq and its partner protein Crc on control regions of different P. aeruginosa target mRNAs. Our results show that these assemblies have mRNA-specific quaternary architectures resulting from the combination of multivalent protein-protein interfaces and recognition of patterns in the RNA sequence. The structural polymorphism of these ribonucleoprotein assemblies enables selective translational repression of many different target mRNAs. This system elucidates how highly complex regulatory pathways can evolve with a minimal economy of proteinogenic components in combination with RNA sequence and fold. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16264.map.gz emd_16264.map.gz | 7.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16264-v30.xml emd-16264-v30.xml emd-16264.xml emd-16264.xml | 17.5 KB 17.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_16264.png emd_16264.png | 66.5 KB | ||
| Others |  emd_16264_half_map_1.map.gz emd_16264_half_map_1.map.gz emd_16264_half_map_2.map.gz emd_16264_half_map_2.map.gz | 95.7 MB 95.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16264 http://ftp.pdbj.org/pub/emdb/structures/EMD-16264 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16264 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16264 | HTTPS FTP |
-Related structure data
| Related structure data |  8bvhMC  8bvjC  8bvmC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_16264.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16264.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Locally filtered cryo-EM map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half map 1
| File | emd_16264_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
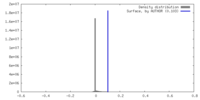
| Density Histograms |
-Half map: Half map 2
| File | emd_16264_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
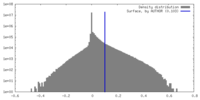
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of RNA chaperone Hfq and helper protein Crc assembled on ...
| Entire | Name: Complex of RNA chaperone Hfq and helper protein Crc assembled on a fragment of amiE mRNA |
|---|---|
| Components |
|
-Supramolecule #1: Complex of RNA chaperone Hfq and helper protein Crc assembled on ...
| Supramolecule | Name: Complex of RNA chaperone Hfq and helper protein Crc assembled on a fragment of amiE mRNA type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 260 KDa |
-Macromolecule #1: RNA-binding protein Hfq
| Macromolecule | Name: RNA-binding protein Hfq / type: protein_or_peptide / ID: 1 / Number of copies: 18 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 9.114487 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSKGHSLQDP YLNTLRKERV PVSIYLVNGI KLQGQIESFD QFVILLKNTV SQMVYKHAIS TVVPSRPVRL PSGDQPAEPG NA |
-Macromolecule #2: Catabolite repression control protein
| Macromolecule | Name: Catabolite repression control protein / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO / EC number: exodeoxyribonuclease III |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 30.101869 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GPAMRIISVN VNGIQAAAER GLLSWLQAQN ADVICLQDTR ASAFDLDDPS FQLDGYFLYA CDAELPEQGG VALYSRLQPK AVISGLGFE TADRYGRYLQ ADFDKVSIAT LLLPSGQSGD ESLNQKFKFM DDFTHYLSKQ RRKRREYIYC GSLYVAHQKM D VKNWRECQ ...String: GPAMRIISVN VNGIQAAAER GLLSWLQAQN ADVICLQDTR ASAFDLDDPS FQLDGYFLYA CDAELPEQGG VALYSRLQPK AVISGLGFE TADRYGRYLQ ADFDKVSIAT LLLPSGQSGD ESLNQKFKFM DDFTHYLSKQ RRKRREYIYC GSLYVAHQKM D VKNWRECQ QMPGFLAPER AWLDEVFGNL GYADALREVS REGDQFSWWP DSEQAEMLNL GWRFDYQVLT PGLRRFVRNA KL PRQPRFS QHAPLIVDYD WQLSI |
-Macromolecule #3: amiE
| Macromolecule | Name: amiE / type: rna / ID: 3 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 33.720895 KDa |
| Sequence | String: UUUUUUCGUC CCGAAAAAAU AACAACAAGA GGUGAUAUCC AUGCGUCACG GCGAUAUU(N)(N) (N)(N)(N)(N)(N) (N)UCC AGCAGCAACG ACACCGUCGG AGUGGCGGUG GUCAACUAC |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: Quantifoil / Material: COPPER / Support film - Material: GRAPHENE OXIDE / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.6 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.6 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 63000 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: cryoSPARC (ver. 2.15) |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: cryoSPARC (ver. 2.15) |
-Atomic model buiding 1
| Refinement | Protocol: RIGID BODY FIT / Target criteria: Cross-correlation coefficient |
|---|---|
| Output model |  PDB-8bvh: |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)