[English] 日本語
 Yorodumi
Yorodumi- EMDB-15835: Cryo-EM structure of homomeric LRRC8C Volume-Regulated Anion Channel -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of homomeric LRRC8C Volume-Regulated Anion Channel | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Ion channel / Volume-regulated anion channel / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationMiscellaneous transport and binding events / volume-sensitive anion channel activity / aspartate transmembrane transport / cyclic-GMP-AMP transmembrane import across plasma membrane / taurine transmembrane transport / monoatomic anion transmembrane transport / cellular response to osmotic stress / protein hexamerization / monoatomic ion channel complex / fat cell differentiation ...Miscellaneous transport and binding events / volume-sensitive anion channel activity / aspartate transmembrane transport / cyclic-GMP-AMP transmembrane import across plasma membrane / taurine transmembrane transport / monoatomic anion transmembrane transport / cellular response to osmotic stress / protein hexamerization / monoatomic ion channel complex / fat cell differentiation / endoplasmic reticulum membrane / endoplasmic reticulum / plasma membrane / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
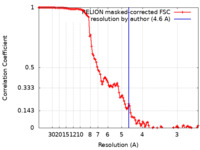
| Method | single particle reconstruction / cryo EM / Resolution: 4.6 Å | |||||||||
 Authors Authors | Sawicka M / Dutzler R | |||||||||
| Funding support |  Switzerland, 1 items Switzerland, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2023 Journal: Nat Struct Mol Biol / Year: 2023Title: Structure of a volume-regulated heteromeric LRRC8A/C channel. Authors: Sonja Rutz / Dawid Deneka / Antje Dittmann / Marta Sawicka / Raimund Dutzler /  Abstract: Volume-regulated anion channels (VRACs) participate in the cellular response to osmotic swelling. These membrane proteins consist of heteromeric assemblies of LRRC8 subunits, whose compositions ...Volume-regulated anion channels (VRACs) participate in the cellular response to osmotic swelling. These membrane proteins consist of heteromeric assemblies of LRRC8 subunits, whose compositions determine permeation properties. Although structures of the obligatory LRRC8A, also referred to as SWELL1, have previously defined the architecture of VRACs, the organization of heteromeric channels has remained elusive. Here we have addressed this question by the structural characterization of murine LRRC8A/C channels. Like LRRC8A, these proteins assemble as hexamers. Despite 12 possible arrangements, we find a predominant organization with an A:C ratio of two. In this assembly, four LRRC8A subunits cluster in their preferred conformation observed in homomers, as pairs of closely interacting proteins that stabilize a closed state of the channel. In contrast, the two interacting LRRC8C subunits show a larger flexibility, underlining their role in the destabilization of the tightly packed A subunits, thereby enhancing the activation properties of the protein. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15835.map.gz emd_15835.map.gz | 11.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15835-v30.xml emd-15835-v30.xml emd-15835.xml emd-15835.xml | 21.1 KB 21.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15835_fsc.xml emd_15835_fsc.xml | 11.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_15835.png emd_15835.png | 77.2 KB | ||
| Filedesc metadata |  emd-15835.cif.gz emd-15835.cif.gz | 6.6 KB | ||
| Others |  emd_15835_additional_1.map.gz emd_15835_additional_1.map.gz emd_15835_half_map_1.map.gz emd_15835_half_map_1.map.gz emd_15835_half_map_2.map.gz emd_15835_half_map_2.map.gz | 8.4 MB 113.8 MB 113.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15835 http://ftp.pdbj.org/pub/emdb/structures/EMD-15835 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15835 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15835 | HTTPS FTP |
-Related structure data
| Related structure data |  8b40MC  8b41C  8b42C  8benC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_15835.map.gz / Format: CCP4 / Size: 144.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15835.map.gz / Format: CCP4 / Size: 144.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.302 Å | ||||||||||||||||||||||||||||||||||||
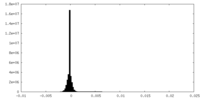
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Masked transmembrane region at 4.1 A
| File | emd_15835_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Masked transmembrane region at 4.1 A | ||||||||||||
| Projections & Slices |
| ||||||||||||
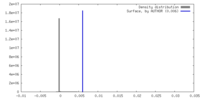
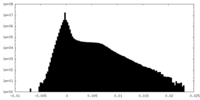
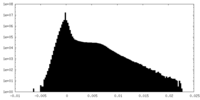
| Density Histograms |
-Half map: #2
| File | emd_15835_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
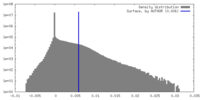
| Density Histograms |
-Half map: #1
| File | emd_15835_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
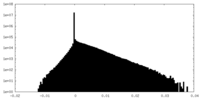
| Density Histograms |
- Sample components
Sample components
-Entire : Homomeric LRRC8C Volume-Regulated Anion Channel
| Entire | Name: Homomeric LRRC8C Volume-Regulated Anion Channel |
|---|---|
| Components |
|
-Supramolecule #1: Homomeric LRRC8C Volume-Regulated Anion Channel
| Supramolecule | Name: Homomeric LRRC8C Volume-Regulated Anion Channel / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Volume-regulated anion channel subunit LRRC8C
| Macromolecule | Name: Volume-regulated anion channel subunit LRRC8C / type: protein_or_peptide / ID: 1 / Number of copies: 7 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 93.472594 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MSIPVTEFRQ FSEQQPAFRV LKPWWDVFTD YLSVAMLMIG VFGCTLQVMQ DKIICLPKRV QPAQNHSSVP NVSQAVISTT PLPPPKPSP TNPATVEMKG LKTDLDLQQY SFINQMCYER ALHWYAKYFP YLVLIHTLVF MLCSNFWFKF PGSSSKIEHF I SILGKCFD ...String: MSIPVTEFRQ FSEQQPAFRV LKPWWDVFTD YLSVAMLMIG VFGCTLQVMQ DKIICLPKRV QPAQNHSSVP NVSQAVISTT PLPPPKPSP TNPATVEMKG LKTDLDLQQY SFINQMCYER ALHWYAKYFP YLVLIHTLVF MLCSNFWFKF PGSSSKIEHF I SILGKCFD SPWTTRALSE VSGEDSEEKD NRKNNMNRSG TIQSGPEGNL VRSQSLKSIP EKFVVDKSAA GALDKKEGEQ AK ALFEKVK KFRLHVEEGD ILYAMYVRQT VLKVIKFLII IAYNSALVSK VQFTVDCNVD IQDMTGYKNF SCNHTMAHLF SKL SFCYLC FVSIYGLTCL YTLYWLFYRS LREYSFEYVR QETGIDDIPD VKNDFAFMLH MIDQYDPLYS KRFAVFLSEV SENK LKQLN LNNEWTPDKL RQKLQTNAHN RLELPLIMLS GLPDTVFEIT ELQSLKLEII KNVMIPATIA QLDNLQELCL HQCSV KIHS AALSFLKENL KVLSVKFDDM RELPPWMYGL RNLEELYLVG SLSHDISKNV TLESLRDLKS LKILSIKSNV SKIPQA VVD VSSHLQKMCV HNDGTKLVML NNLKKMTNLT ELELVHCDLE RIPHAVFSLL SLQELDLKEN NLKSIEEIVS FQHLRKL TV LKLWYNSIAY IPEHIKKLTS LERLFFSHNK VEVLPSHLFL CNKIRYLDLS YNDIRFIPPE IGVLQSLQYF SITCNKVE S LPDELYFCKK LKTLKIGKNS LSVLSPKIGN LLFLSYLDIK GNHFEVLPPE LGDCRALKRA RLVVEDALFE TLPSDVREQ MKADALEVLF Q UniProtKB: Volume-regulated anion channel subunit LRRC8C |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE-PROPANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.4 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller


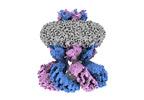











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)