[English] 日本語
 Yorodumi
Yorodumi- EMDB-15578: CryoEM structure of the Chikungunya virus nsP1 capping pores in c... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of the Chikungunya virus nsP1 capping pores in covalent complex with a 7GMP cap structure | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Alphavirus Replication complex Capping pores Membrane pore Methyltransferase gunayltransferase VIRAL PROTEIN / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationADP-ribose 1''-phosphate phosphatase / host cell filopodium / mRNA methyltransferase activity / mRNA 5'-phosphatase / mRNA 5'-triphosphate monophosphatase activity / polynucleotide adenylyltransferase / poly(A) RNA polymerase activity / mRNA modification / regulation of cytoskeleton organization / symbiont-mediated suppression of host mRNA transcription via inhibition of RNA polymerase II activity ...ADP-ribose 1''-phosphate phosphatase / host cell filopodium / mRNA methyltransferase activity / mRNA 5'-phosphatase / mRNA 5'-triphosphate monophosphatase activity / polynucleotide adenylyltransferase / poly(A) RNA polymerase activity / mRNA modification / regulation of cytoskeleton organization / symbiont-mediated suppression of host mRNA transcription via inhibition of RNA polymerase II activity / 7-methylguanosine mRNA capping / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT1 activity / cysteine-type peptidase activity / Transferases; Transferring one-carbon groups; Methyltransferases / host cell cytoplasmic vesicle membrane / Transferases; Transferring phosphorus-containing groups; Nucleotidyltransferases / nucleoside-triphosphate phosphatase / methylation / Hydrolases; Acting on peptide bonds (peptidases); Cysteine endopeptidases / RNA helicase activity / symbiont-mediated suppression of host innate immune response / RNA helicase / symbiont-mediated suppression of host type I interferon-mediated signaling pathway / symbiont-mediated suppression of host gene expression / RNA-directed RNA polymerase / viral RNA genome replication / RNA-directed RNA polymerase activity / DNA-templated transcription / GTP binding / host cell plasma membrane / host cell nucleus / ATP hydrolysis activity / proteolysis / RNA binding / ATP binding / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Chikungunya virus strain S27-African prototype Chikungunya virus strain S27-African prototype | |||||||||
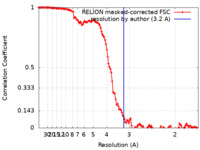
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||
 Authors Authors | Jones R / Hons M / Reguera J | |||||||||
| Funding support |  France, 1 items France, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2023 Journal: Proc Natl Acad Sci U S A / Year: 2023Title: Structural basis and dynamics of Chikungunya alphavirus RNA capping by nsP1 capping pores. Authors: Rhian Jones / Michael Hons / Nadia Rabah / Noelia Zamarreño / Rocío Arranz / Juan Reguera /   Abstract: Alphaviruses are emerging positive-stranded RNA viruses which replicate and transcribe their genomes in membranous organelles formed in the cell cytoplasm. The nonstructural protein 1 (nsP1) is ...Alphaviruses are emerging positive-stranded RNA viruses which replicate and transcribe their genomes in membranous organelles formed in the cell cytoplasm. The nonstructural protein 1 (nsP1) is responsible for viral RNA capping and gates the replication organelles by assembling into monotopic membrane-associated dodecameric pores. The capping pathway is unique to Alphaviruses; beginning with the N methylation of a guanosine triphosphate (GTP) molecule, followed by the covalent linkage of an mGMP group to a conserved histidine in nsP1 and the transfer of this cap structure to a diphosphate RNA. Here, we provide structural snapshots of different stages of the reaction pathway showing how nsP1 pores recognize the substrates of the methyl-transfer reaction, GTP and S-adenosyl methionine (SAM), how the enzyme reaches a metastable postmethylation state with SAH and mGTP in the active site, and the subsequent covalent transfer of mGMP to nsP1 triggered by the presence of RNA and postdecapping reaction conformational changes inducing the opening of the pore. In addition, we biochemically characterize the capping reaction, demonstrating specificity for the RNA substrate and the reversibility of the cap transfer resulting in decapping activity and the release of reaction intermediates. Our data identify the molecular determinants allowing each pathway transition, providing an explanation for the need for the SAM methyl donor all along the pathway and clues about the conformational rearrangements associated to the enzymatic activity of nsP1. Together, our results set ground for the structural and functional understanding of alphavirus RNA-capping and the design of antivirals. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15578.map.gz emd_15578.map.gz | 157.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15578-v30.xml emd-15578-v30.xml emd-15578.xml emd-15578.xml | 24 KB 24 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15578_fsc.xml emd_15578_fsc.xml | 12.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_15578.png emd_15578.png | 97.1 KB | ||
| Filedesc metadata |  emd-15578.cif.gz emd-15578.cif.gz | 7.3 KB | ||
| Others |  emd_15578_half_map_1.map.gz emd_15578_half_map_1.map.gz emd_15578_half_map_2.map.gz emd_15578_half_map_2.map.gz | 164.9 MB 165 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15578 http://ftp.pdbj.org/pub/emdb/structures/EMD-15578 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15578 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15578 | HTTPS FTP |
-Related structure data
| Related structure data |  8apxMC  8aovC  8aowC  8aoxC  8axvC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_15578.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15578.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.855 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_15578_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
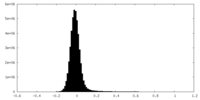
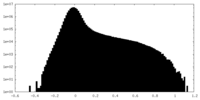
| Density Histograms |
-Half map: #1
| File | emd_15578_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : CHIKV nsP1 capping pores covalently linked to m7GMP cap via H37
| Entire | Name: CHIKV nsP1 capping pores covalently linked to m7GMP cap via H37 |
|---|---|
| Components |
|
-Supramolecule #1: CHIKV nsP1 capping pores covalently linked to m7GMP cap via H37
| Supramolecule | Name: CHIKV nsP1 capping pores covalently linked to m7GMP cap via H37 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Chikungunya virus strain S27-African prototype Chikungunya virus strain S27-African prototype |
| Molecular weight | Theoretical: 720 KDa |
-Macromolecule #1: Polyprotein P1234
| Macromolecule | Name: Polyprotein P1234 / type: protein_or_peptide / ID: 1 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Chikungunya virus strain S27-African prototype / Strain: S27-African prototype Chikungunya virus strain S27-African prototype / Strain: S27-African prototype |
| Molecular weight | Theoretical: 53.011754 KDa |
| Recombinant expression | Organism:  Baculovirus expression vector pFastBac1-HM Baculovirus expression vector pFastBac1-HM |
| Sequence | String: MDPVYVDIDA DSAFLKALQR AYPMFEVEPR QVTPND(PJ3)ANA RAFSHLAIKL IEQEIDPDST ILDIGSAPAR RMMSDR KYH CVCPMRSAED PERLANYARK LASAAGKVLD RNISGKIGDL QAVMAVPDTE TPTFCLHTDV SCRQRADVAI YQDVYAV HA PTSLYHQAIK ...String: MDPVYVDIDA DSAFLKALQR AYPMFEVEPR QVTPND(PJ3)ANA RAFSHLAIKL IEQEIDPDST ILDIGSAPAR RMMSDR KYH CVCPMRSAED PERLANYARK LASAAGKVLD RNISGKIGDL QAVMAVPDTE TPTFCLHTDV SCRQRADVAI YQDVYAV HA PTSLYHQAIK GVRLAYWVGF DTTPFMYNAM AGAYPSYSTN WADEQVLKAK NIGLCSTDLT EGRRGKLSIM RGKKLEPC D RVLFSVGSTL YPESRKLLKS WHLPSVFHLK GKLSFTCRCD TVVSCEGYVV KRITMSPGLY GKTTGYAVTH HADGFLMCK TTDTVDGERV SFSVCTYVPA TICDQMTGIL ATEVTPEDAQ KLLVGLNQRI VVNGRTQRNT NTMKNYMIPV VAQAFSKWAK ECRKDMEDE KLLGVRERTL TCCCLWAFKK QKTHTVYKRP DTQSIQKVQA EFDSFVVPSL WSSGLSIPLR TRIK UniProtKB: Polyprotein P1234 |
-Macromolecule #2: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 2 / Number of copies: 12 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Macromolecule #3: S-ADENOSYL-L-HOMOCYSTEINE
| Macromolecule | Name: S-ADENOSYL-L-HOMOCYSTEINE / type: ligand / ID: 3 / Number of copies: 12 / Formula: SAH |
|---|---|
| Molecular weight | Theoretical: 384.411 Da |
| Chemical component information |  ChemComp-SAH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.2 mg/mL | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.6 Component:
| ||||||||||||||||||
| Grid | Model: Quantifoil R2/2 / Material: GOLD / Mesh: 300 / Support film - Material: GRAPHENE OXIDE / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 10 sec. / Pretreatment - Atmosphere: AIR | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 298 K / Instrument: FEI VITROBOT MARK IV | ||||||||||||||||||
| Details | Sample incubated with 10x molar excess of m7GTP, SAH and 27mer CHIKV RNA to form the complex prior to freezing in the presence of magnesium. |
- Electron microscopy
Electron microscopy
| Microscope | TFS TALOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 773 / Average exposure time: 38.0 sec. / Average electron dose: 32.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.8 µm |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Target criteria: Correlation coefficient |
| Output model |  PDB-8apx: |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)