[English] 日本語
 Yorodumi
Yorodumi- EMDB-14082: Structure of the 26S proteasome-Ubp6 complex in the si state (Cor... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the 26S proteasome-Ubp6 complex in the si state (Core Particle and Lid) | ||||||||||||||||||
 Map data Map data | |||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | Proteasome UBP6 Ubiquitin Allostery / MOTOR PROTEIN | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationSAGA complex localization to transcription regulatory region / Metalloprotease DUBs / proteasome storage granule assembly / transcription export complex 2 / protein deneddylation / peroxisome fission / maintenance of DNA trinucleotide repeats / filamentous growth / COP9 signalosome / proteasome regulatory particle ...SAGA complex localization to transcription regulatory region / Metalloprotease DUBs / proteasome storage granule assembly / transcription export complex 2 / protein deneddylation / peroxisome fission / maintenance of DNA trinucleotide repeats / filamentous growth / COP9 signalosome / proteasome regulatory particle / mitochondrial fission / proteasome regulatory particle, lid subcomplex / proteasome regulatory particle, base subcomplex / metal-dependent deubiquitinase activity / K48-linked polyubiquitin modification-dependent protein binding / proteasome core complex assembly / nuclear outer membrane-endoplasmic reticulum membrane network / Proteasome assembly / Cross-presentation of soluble exogenous antigens (endosomes) / TNFR2 non-canonical NF-kB pathway / proteasomal ubiquitin-independent protein catabolic process / Ubiquitin-Mediated Degradation of Phosphorylated Cdc25A / Regulation of PTEN stability and activity / CDK-mediated phosphorylation and removal of Cdc6 / FBXL7 down-regulates AURKA during mitotic entry and in early mitosis / KEAP1-NFE2L2 pathway / Neddylation / proteasome binding / Orc1 removal from chromatin / MAPK6/MAPK4 signaling / regulation of protein catabolic process / proteasome storage granule / Antigen processing: Ubiquitination & Proteasome degradation / polyubiquitin modification-dependent protein binding / protein deubiquitination / proteasome endopeptidase complex / Ub-specific processing proteases / endopeptidase activator activity / proteasome core complex, beta-subunit complex / proteasome assembly / threonine-type endopeptidase activity / proteasome core complex, alpha-subunit complex / mRNA export from nucleus / enzyme regulator activity / Neutrophil degranulation / protein folding chaperone / proteasome complex / ubiquitin binding / double-strand break repair via homologous recombination / metallopeptidase activity / peroxisome / ubiquitin-dependent protein catabolic process / protein-macromolecule adaptor activity / endopeptidase activity / molecular adaptor activity / proteasome-mediated ubiquitin-dependent protein catabolic process / ubiquitinyl hydrolase 1 / cysteine-type deubiquitinase activity / regulation of cell cycle / mRNA binding / ubiquitin protein ligase binding / endoplasmic reticulum membrane / structural molecule activity / endoplasmic reticulum / positive regulation of transcription by RNA polymerase II / mitochondrion / metal ion binding / nucleus / cytosol / cytoplasm Similarity search - Function | ||||||||||||||||||
| Biological species |  | ||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 6.1 Å | ||||||||||||||||||
 Authors Authors | Hung KYS / Klumpe S | ||||||||||||||||||
| Funding support |  United States, 5 items United States, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Allosteric control of Ubp6 and the proteasome via a bidirectional switch. Authors: Ka Ying Sharon Hung / Sven Klumpe / Markus R Eisele / Suzanne Elsasser / Geng Tian / Shuangwu Sun / Jamie A Moroco / Tat Cheung Cheng / Tapan Joshi / Timo Seibel / Duco Van Dalen / Xin-Hua ...Authors: Ka Ying Sharon Hung / Sven Klumpe / Markus R Eisele / Suzanne Elsasser / Geng Tian / Shuangwu Sun / Jamie A Moroco / Tat Cheung Cheng / Tapan Joshi / Timo Seibel / Duco Van Dalen / Xin-Hua Feng / Ying Lu / Huib Ovaa / John R Engen / Byung-Hoon Lee / Till Rudack / Eri Sakata / Daniel Finley /      Abstract: The proteasome recognizes ubiquitinated proteins and can also edit ubiquitin marks, allowing substrates to be rejected based on ubiquitin chain topology. In yeast, editing is mediated by ...The proteasome recognizes ubiquitinated proteins and can also edit ubiquitin marks, allowing substrates to be rejected based on ubiquitin chain topology. In yeast, editing is mediated by deubiquitinating enzyme Ubp6. The proteasome activates Ubp6, whereas Ubp6 inhibits the proteasome through deubiquitination and a noncatalytic effect. Here, we report cryo-EM structures of the proteasome bound to Ubp6, based on which we identify mutants in Ubp6 and proteasome subunit Rpt1 that abrogate Ubp6 activation. The Ubp6 mutations define a conserved region that we term the ILR element. The ILR is found within the BL1 loop, which obstructs the catalytic groove in free Ubp6. Rpt1-ILR interaction opens the groove by rearranging not only BL1 but also a previously undescribed network of three interconnected active-site-blocking loops. Ubp6 activation and noncatalytic proteasome inhibition are linked in that they are eliminated by the same mutations. Ubp6 and ubiquitin together drive proteasomes into a unique conformation associated with proteasome inhibition. Thus, a multicomponent allosteric switch exerts simultaneous control over both Ubp6 and the proteasome. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14082.map.gz emd_14082.map.gz | 55.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14082-v30.xml emd-14082-v30.xml emd-14082.xml emd-14082.xml | 42.8 KB 42.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_14082.png emd_14082.png | 116.3 KB | ||
| Filedesc metadata |  emd-14082.cif.gz emd-14082.cif.gz | 11.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14082 http://ftp.pdbj.org/pub/emdb/structures/EMD-14082 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14082 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14082 | HTTPS FTP |
-Validation report
| Summary document |  emd_14082_validation.pdf.gz emd_14082_validation.pdf.gz | 557.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_14082_full_validation.pdf.gz emd_14082_full_validation.pdf.gz | 557.3 KB | Display | |
| Data in XML |  emd_14082_validation.xml.gz emd_14082_validation.xml.gz | 6.6 KB | Display | |
| Data in CIF |  emd_14082_validation.cif.gz emd_14082_validation.cif.gz | 7.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14082 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14082 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14082 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14082 | HTTPS FTP |
-Related structure data
| Related structure data |  7qo3MC  7qo5C  7qo6C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_14082.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14082.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.1 Å | ||||||||||||||||||||||||||||||||||||
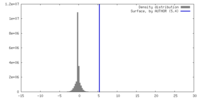
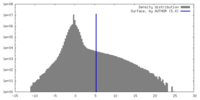
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
+Entire : 26S proteasome-Ubp6 complex (si state)
+Supramolecule #1: 26S proteasome-Ubp6 complex (si state)
+Macromolecule #1: Proteasome subunit alpha type-1
+Macromolecule #2: Proteasome subunit alpha type-2
+Macromolecule #3: Proteasome subunit alpha type-3
+Macromolecule #4: Proteasome subunit alpha type-4
+Macromolecule #5: Proteasome subunit alpha type-5
+Macromolecule #6: Proteasome subunit alpha type-6
+Macromolecule #7: Probable proteasome subunit alpha type-7
+Macromolecule #8: Proteasome subunit beta type-1
+Macromolecule #9: Proteasome subunit beta type-2
+Macromolecule #10: Proteasome subunit beta type-3
+Macromolecule #11: Proteasome subunit beta type-4
+Macromolecule #12: Proteasome subunit beta type-5
+Macromolecule #13: Proteasome subunit beta type-6
+Macromolecule #14: Proteasome subunit beta type-7
+Macromolecule #15: 26S proteasome regulatory subunit RPN10
+Macromolecule #16: Ubiquitin carboxyl-terminal hydrolase RPN11
+Macromolecule #17: 26S proteasome regulatory subunit RPN12
+Macromolecule #18: 26S proteasome regulatory subunit RPN13
+Macromolecule #19: 26S proteasome complex subunit SEM1
+Macromolecule #20: 26S proteasome regulatory subunit RPN1
+Macromolecule #21: 26S proteasome regulatory subunit RPN2
+Macromolecule #22: 26S proteasome regulatory subunit RPN3
+Macromolecule #23: 26S proteasome regulatory subunit RPN5
+Macromolecule #24: 26S proteasome regulatory subunit RPN6
+Macromolecule #25: 26S proteasome regulatory subunit RPN7
+Macromolecule #26: 26S proteasome regulatory subunit RPN8
+Macromolecule #27: 26S proteasome regulatory subunit RPN9
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: EMDB MAP |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 6.1 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 88243 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller

























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)