[English] 日本語
 Yorodumi
Yorodumi- EMDB-13784: Cryo-EM structure of clamped S.cerevisiae condensin-DNA complex (... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
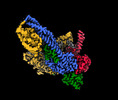
| Title | Cryo-EM structure of clamped S.cerevisiae condensin-DNA complex (form II) | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | condensin / SMC / CELL CYCLE | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of meiotic DNA double-strand break formation / tRNA gene clustering / Condensation of Prometaphase Chromosomes / meiotic chromosome condensation / meiotic chromosome separation / condensin complex / DNA secondary structure binding / maintenance of rDNA / rDNA chromatin condensation / nucleophagy ...negative regulation of meiotic DNA double-strand break formation / tRNA gene clustering / Condensation of Prometaphase Chromosomes / meiotic chromosome condensation / meiotic chromosome separation / condensin complex / DNA secondary structure binding / maintenance of rDNA / rDNA chromatin condensation / nucleophagy / synaptonemal complex assembly / condensed chromosome, centromeric region / mitotic chromosome condensation / chromosome condensation / silent mating-type cassette heterochromatin formation / minor groove of adenine-thymine-rich DNA binding / mitotic sister chromatid segregation / condensed chromosome / double-stranded DNA binding / histone binding / cell division / chromatin binding / chromatin / nucleolus / ATP hydrolysis activity / mitochondrion / ATP binding / nucleus / cytoplasm Similarity search - Function | ||||||||||||
| Biological species |  | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.0 Å | ||||||||||||
 Authors Authors | Lee B-G / Rhodes J / Lowe J | ||||||||||||
| Funding support |  United Kingdom, 3 items United Kingdom, 3 items
| ||||||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2022 Journal: Proc Natl Acad Sci U S A / Year: 2022Title: Clamping of DNA shuts the condensin neck gate. Authors: Byung-Gil Lee / James Rhodes / Jan Löwe /  Abstract: SignificanceDNA needs to be compacted to fit into nuclei and during cell division, when dense chromatids are formed for their mechanical segregation, a process that depends on the protein complex ...SignificanceDNA needs to be compacted to fit into nuclei and during cell division, when dense chromatids are formed for their mechanical segregation, a process that depends on the protein complex condensin. It forms and enlarges loops in DNA through loop extrusion. Our work resolves the atomic structure of a DNA-bound state of condensin in which ATP has not been hydrolyzed. The DNA is clamped within a compartment that has been reported previously in other structural maintenance of chromosomes (SMC) complexes, including Rad50, cohesin, and MukBEF. With the caveat of important differences, it means that all SMC complexes cycle through at least some similar states and undergo similar conformational changes in their head modules, while hydrolyzing ATP and translocating DNA. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13784.map.gz emd_13784.map.gz | 13.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13784-v30.xml emd-13784-v30.xml emd-13784.xml emd-13784.xml | 22.4 KB 22.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_13784.png emd_13784.png | 81 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13784 http://ftp.pdbj.org/pub/emdb/structures/EMD-13784 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13784 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13784 | HTTPS FTP |
-Related structure data
| Related structure data |  7q2yMC  7q2xC  7q2zC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_13784.map.gz / Format: CCP4 / Size: 16.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13784.map.gz / Format: CCP4 / Size: 16.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
+Entire : complex of condensin tetramer (smc2, smc4, brn1, ycs4), DNA, ADP-BeF3
+Supramolecule #1: complex of condensin tetramer (smc2, smc4, brn1, ycs4), DNA, ADP-BeF3
+Macromolecule #1: Structural maintenance of chromosomes protein 2
+Macromolecule #2: Structural maintenance of chromosomes protein 4
+Macromolecule #3: Condensin complex subunit 2
+Macromolecule #4: Condensin complex subunit 1
+Macromolecule #5: DNA (28-MER)
+Macromolecule #6: DNA (28-MER)
+Macromolecule #7: ADENOSINE-5'-DIPHOSPHATE
+Macromolecule #8: BERYLLIUM TRIFLUORIDE ION
+Macromolecule #9: MAGNESIUM ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 55.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.0 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION / Number images used: 251999 |
| Initial angle assignment | Type: ANGULAR RECONSTITUTION / Software - Name: RELION |
| Final angle assignment | Type: ANGULAR RECONSTITUTION / Software - Name: RELION |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)
























