[English] 日本語
 Yorodumi
Yorodumi- EMDB-12886: Structure of the apo-state of the bacteriophage PhiKZ non-virion ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12886 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the apo-state of the bacteriophage PhiKZ non-virion RNA polymerase | |||||||||
 Map data Map data | Structure of the apo-state of the bacteriophage PhiKZ non-virion RNA polymerase | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | RNA polymerase / PhiKZ / non-virion / TRANSCRIPTION | |||||||||
| Function / homology | PHIKZ123 / PHIKZ074 / PHIKZ068 / PHIKZ055 Function and homology information Function and homology information | |||||||||
| Biological species |  Pseudomonas virus phiKZ / Pseudomonas virus phiKZ /  Pseudomonas phage PA7 (virus) / Pseudomonas phage PA7 (virus) /  Pseudomonas phage phiKZ (virus) Pseudomonas phage phiKZ (virus) | |||||||||
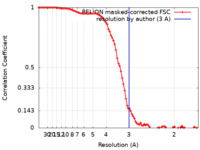
| Method | single particle reconstruction / cryo EM / Resolution: 3.0 Å | |||||||||
 Authors Authors | de Martin Garrido N / Lai Wan Loong YTE | |||||||||
| Funding support |  United Kingdom, United Kingdom,  Russian Federation, 2 items Russian Federation, 2 items
| |||||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2021 Journal: Nucleic Acids Res / Year: 2021Title: Structure of the bacteriophage PhiKZ non-virion RNA polymerase. Authors: Natàlia deYMartín Garrido / Mariia Orekhova / Yuen Ting Emilie Lai Wan Loong / Anna Litvinova / Kailash Ramlaul / Tatyana Artamonova / Alexei S Melnikov / Pavel Serdobintsev / Christopher ...Authors: Natàlia deYMartín Garrido / Mariia Orekhova / Yuen Ting Emilie Lai Wan Loong / Anna Litvinova / Kailash Ramlaul / Tatyana Artamonova / Alexei S Melnikov / Pavel Serdobintsev / Christopher H S Aylett / Maria Yakunina /   Abstract: Bacteriophage ΦKZ (PhiKZ) is the archetype of a family of massive bacterial viruses. It is considered to have therapeutic potential as its host, Pseudomonas aeruginosa, is an opportunistic, ...Bacteriophage ΦKZ (PhiKZ) is the archetype of a family of massive bacterial viruses. It is considered to have therapeutic potential as its host, Pseudomonas aeruginosa, is an opportunistic, intrinsically antibiotic resistant, pathogen that kills tens of thousands worldwide each year. ΦKZ is an incredibly interesting virus, expressing many systems that the host already possesses. On infection, it forms a 'nucleus', erecting a barrier around its genome to exclude host endonucleases and CRISPR-Cas systems. ΦKZ infection is independent of the host transcriptional apparatus. It expresses two different multi-subunit RNA polymerases (RNAPs): the virion RNAP (vRNAP) is injected with the viral DNA during infection to transcribe early genes, including those encoding the non-virion RNAP (nvRNAP), which transcribes all further genes. ΦKZ nvRNAP is formed by four polypeptides thought to represent homologues of the eubacterial β/β' subunits, and a fifth with unclear homology, but essential for transcription. We have resolved the structure of ΦKZ nvRNAP to better than 3.0 Å, shedding light on its assembly, homology, and the biological role of the fifth subunit: it is an embedded, integral member of the complex, the position, structural homology and biochemical role of which imply that it has evolved from an ancestral homologue to σ-factor. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12886.map.gz emd_12886.map.gz | 59.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12886-v30.xml emd-12886-v30.xml emd-12886.xml emd-12886.xml | 29.3 KB 29.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_12886_fsc.xml emd_12886_fsc.xml | 9.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_12886.png emd_12886.png | 131.8 KB | ||
| Filedesc metadata |  emd-12886.cif.gz emd-12886.cif.gz | 8.5 KB | ||
| Others |  emd_12886_half_map_1.map.gz emd_12886_half_map_1.map.gz emd_12886_half_map_2.map.gz emd_12886_half_map_2.map.gz | 49.6 MB 49.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12886 http://ftp.pdbj.org/pub/emdb/structures/EMD-12886 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12886 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12886 | HTTPS FTP |
-Related structure data
| Related structure data |  7ogrMC  7ogpC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_12886.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12886.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
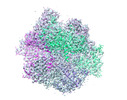
| Annotation | Structure of the apo-state of the bacteriophage PhiKZ non-virion RNA polymerase | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.85 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
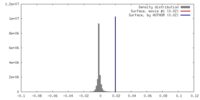
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: Structure of the apo-state of the bacteriophage PhiKZ...
| File | emd_12886_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Structure of the apo-state of the bacteriophage PhiKZ non-virion RNA polymerase - halfmap 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
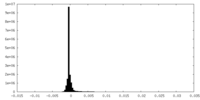
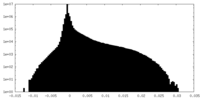
| Density Histograms |
-Half map: Structure of the apo-state of the bacteriophage PhiKZ...
| File | emd_12886_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Structure of the apo-state of the bacteriophage PhiKZ non-virion RNA polymerase - halfmap 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
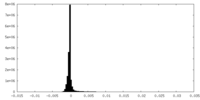
| Density Histograms |
- Sample components
Sample components
+Entire : Bacteriophage PhiKZ non-virion RNA polymerase
+Supramolecule #1: Bacteriophage PhiKZ non-virion RNA polymerase
+Supramolecule #2: Bacteriophage PhiKZ non-virion RNA polymerase
+Supramolecule #3: Bacteriophage PhiKZ non-virion RNA polymerase
+Supramolecule #4: Bacteriophage PhiKZ non-virion RNA polymerase
+Macromolecule #1: UNK helices
+Macromolecule #2: PHIKZ055
+Macromolecule #3: PHIKZ068
+Macromolecule #4: DNA-directed RNA polymerase
+Macromolecule #5: PHIKZ074
+Macromolecule #6: PHIKZ123
+Macromolecule #7: ZINC ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL |
|---|---|
| Buffer | pH: 8 Details: 15 mM Tris-Cl pH 8.0, 150 mM NaCl, 0.5 mM EDTA, 2 mM MgCl2, 1 mM DTT |
| Grid | Model: Quantifoil R2/1 / Support film - Material: GRAPHENE OXIDE / Support film - topology: CONTINUOUS / Support film - Film thickness: 1 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 278 K / Instrument: FEI VITROBOT MARK IV / Details: 1-2 s blot. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 100.0 K / Max: 100.0 K |
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 70 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Digitization - Dimensions - Width: 5096 pixel / Digitization - Dimensions - Height: 4092 pixel / Number grids imaged: 2 / Number real images: 10582 / Average electron dose: 40.0 e/Å2 / Details: TIFF movie mode |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.25 µm / Nominal defocus min: 0.75 µm / Nominal magnification: 48000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Details | Core conserved regions initially identified from comparison with PDB 6EDT - remainder built de novo |
| Refinement | Protocol: OTHER |
| Output model |  PDB-7ogr: |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)