+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8910 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of murine AA amyloid fibril | |||||||||
 Map data Map data | Cryo-EM reconstruction of murine Serum Amyloid A fibrils from diseased mouse spleen using Relion 2.1. The central 30% of the reconstruction is shown. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | AA-amyloidosis / fibril / cross-beta / helical / PROTEIN FIBRIL | |||||||||
| Function / homology |  Function and homology information Function and homology informationresponse to stilbenoid / high-density lipoprotein particle / cytoplasmic microtubule / acute-phase response / G protein-coupled receptor binding Similarity search - Function | |||||||||
| Biological species |  | |||||||||
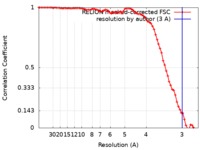
| Method | helical reconstruction / cryo EM / Resolution: 3.0 Å | |||||||||
 Authors Authors | Liberta F / Fandrich M / Schmidt M | |||||||||
| Funding support |  Germany, 2 items Germany, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2019 Journal: Nat Commun / Year: 2019Title: Cryo-EM fibril structures from systemic AA amyloidosis reveal the species complementarity of pathological amyloids. Authors: Falk Liberta / Sarah Loerch / Matthies Rennegarbe / Angelika Schierhorn / Per Westermark / Gunilla T Westermark / Bouke P C Hazenberg / Nikolaus Grigorieff / Marcus Fändrich / Matthias Schmidt /     Abstract: Systemic AA amyloidosis is a worldwide occurring protein misfolding disease of humans and animals. It arises from the formation of amyloid fibrils from the acute phase protein serum amyloid A. Here, ...Systemic AA amyloidosis is a worldwide occurring protein misfolding disease of humans and animals. It arises from the formation of amyloid fibrils from the acute phase protein serum amyloid A. Here, we report the purification and electron cryo-microscopy analysis of amyloid fibrils from a mouse and a human patient with systemic AA amyloidosis. The obtained resolutions are 3.0 Å and 2.7 Å for the murine and human fibril, respectively. The two fibrils differ in fundamental properties, such as presence of right-hand or left-hand twisted cross-β sheets and overall fold of the fibril proteins. Yet, both proteins adopt highly similar β-arch conformations within the N-terminal ~21 residues. Our data demonstrate the importance of the fibril protein N-terminus for the stability of the analyzed amyloid fibril morphologies and suggest strategies of combating this disease by interfering with specific fibril polymorphs. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8910.map.gz emd_8910.map.gz | 2.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8910-v30.xml emd-8910-v30.xml emd-8910.xml emd-8910.xml | 16.4 KB 16.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_8910_fsc.xml emd_8910_fsc.xml | 7.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_8910.png emd_8910.png | 273.8 KB | ||
| Masks |  emd_8910_msk_1.map emd_8910_msk_1.map | 35.3 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-8910.cif.gz emd-8910.cif.gz | 5.6 KB | ||
| Others |  emd_8910_half_map_1.map.gz emd_8910_half_map_1.map.gz emd_8910_half_map_2.map.gz emd_8910_half_map_2.map.gz | 27.2 MB 27.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8910 http://ftp.pdbj.org/pub/emdb/structures/EMD-8910 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8910 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8910 | HTTPS FTP |
-Related structure data
| Related structure data |  6dsoMC  9232C  6mstC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10825 (Title: Cryo electron microscopy of ex-vivo murine SAA amyloid fibrils EMPIAR-10825 (Title: Cryo electron microscopy of ex-vivo murine SAA amyloid fibrilsData size: 533.9 Data #1: Unaligned multiframe micrographs of ex-vivo murine SAA1 [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_8910.map.gz / Format: CCP4 / Size: 35.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8910.map.gz / Format: CCP4 / Size: 35.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM reconstruction of murine Serum Amyloid A fibrils from diseased mouse spleen using Relion 2.1. The central 30% of the reconstruction is shown. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.35 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
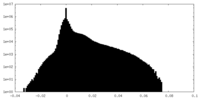
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_8910_msk_1.map emd_8910_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
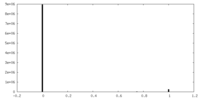
| Density Histograms |
-Half map: Map 2 of two independently refined half maps
| File | emd_8910_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Map 2 of two independently refined half maps | ||||||||||||
| Projections & Slices |
| ||||||||||||
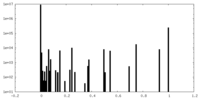
| Density Histograms |
-Half map: Map 1 of two independently refined half maps
| File | emd_8910_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Map 1 of two independently refined half maps | ||||||||||||
| Projections & Slices |
| ||||||||||||
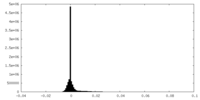
| Density Histograms |
- Sample components
Sample components
-Entire : AA amyloid fibril
| Entire | Name: AA amyloid fibril |
|---|---|
| Components |
|
-Supramolecule #1: AA amyloid fibril
| Supramolecule | Name: AA amyloid fibril / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Serum amyloid A-2 protein
| Macromolecule | Name: Serum amyloid A-2 protein / type: protein_or_peptide / ID: 1 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 9.362094 KDa |
| Sequence | String: GFFSFIGEAF QGAGDMWRAY TDMKEAGWKD GDKYFHARGN YDAAQRGPGG VWAAEKISDA RESFQEFFGR GHEDTMADQE ANR UniProtKB: Serum amyloid A-2 protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | helical array |
- Sample preparation
Sample preparation
| Concentration | 0.2 mg/mL |
|---|---|
| Buffer | pH: 7 / Component - Formula: ddH2O |
| Grid | Model: C-flat-2/1 / Material: COPPER / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 40 sec. / Pretreatment - Atmosphere: OTHER / Details: 20 mA, 0.25 mBar |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 293 K / Instrument: FEI VITROBOT MARK III |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Number real images: 1063 / Average electron dose: 20.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: -5.5 µm / Nominal defocus min: -1.3 µm / Nominal magnification: 105000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-6dso: |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)