+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Caulobacter crescentus FljM flagellar filament (symmetrized) | ||||||||||||||||||
 Map data Map data | FljM (symmetrized) map | ||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | flagellin / flagellar filament / STRUCTURAL PROTEIN | ||||||||||||||||||
| Function / homology | Flagellin, C-terminal domain / Bacterial flagellin C-terminal helical region / Flagellin / Flagellin, N-terminal domain / Bacterial flagellin N-terminal helical region / bacterial-type flagellum / structural molecule activity / extracellular region / Flagellin FljM Function and homology information Function and homology information | ||||||||||||||||||
| Biological species |  Caulobacter vibrioides (bacteria) Caulobacter vibrioides (bacteria) | ||||||||||||||||||
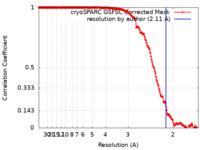
| Method | helical reconstruction / cryo EM / Resolution: 2.11 Å | ||||||||||||||||||
 Authors Authors | Sanchez JC / Montemayor EJ / Ploscariu NT / Parrell D / Baumgardt JK / Yang JE / Sibert B / Cai K / Wright ER | ||||||||||||||||||
| Funding support |  United States, 5 items United States, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Direct evidence for multi-flagellin filament stabilization via atomic-level architecture of Caulobacter crescentus flagellar filaments Authors: Sanchez JC / Montemayor EJ / Ploscariu NT / Parrell D / Baumgardt JK / Yang JE / Sibert B / Cai K / Wright ER | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_42770.map.gz emd_42770.map.gz | 53 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-42770-v30.xml emd-42770-v30.xml emd-42770.xml emd-42770.xml | 17.2 KB 17.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_42770_fsc.xml emd_42770_fsc.xml | 16.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_42770.png emd_42770.png | 114.1 KB | ||
| Masks |  emd_42770_msk_1.map emd_42770_msk_1.map | 512 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-42770.cif.gz emd-42770.cif.gz | 5.9 KB | ||
| Others |  emd_42770_half_map_1.map.gz emd_42770_half_map_1.map.gz emd_42770_half_map_2.map.gz emd_42770_half_map_2.map.gz | 476 MB 476 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-42770 http://ftp.pdbj.org/pub/emdb/structures/EMD-42770 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42770 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42770 | HTTPS FTP |
-Validation report
| Summary document |  emd_42770_validation.pdf.gz emd_42770_validation.pdf.gz | 980.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_42770_full_validation.pdf.gz emd_42770_full_validation.pdf.gz | 980.2 KB | Display | |
| Data in XML |  emd_42770_validation.xml.gz emd_42770_validation.xml.gz | 26.5 KB | Display | |
| Data in CIF |  emd_42770_validation.cif.gz emd_42770_validation.cif.gz | 34.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-42770 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-42770 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-42770 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-42770 | HTTPS FTP |
-Related structure data
| Related structure data |  8uxnMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_42770.map.gz / Format: CCP4 / Size: 476.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_42770.map.gz / Format: CCP4 / Size: 476.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | FljM (symmetrized) map | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.834 Å | ||||||||||||||||||||
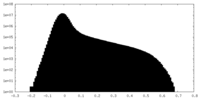
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_42770_msk_1.map emd_42770_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
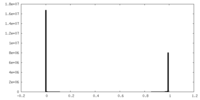
| Density Histograms |
-Half map: FljM (symmetrized) half map 2
| File | emd_42770_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | FljM (symmetrized) half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
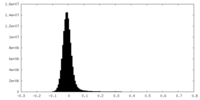
| Density Histograms |
-Half map: FljM (symmetrized) half map 1
| File | emd_42770_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | FljM (symmetrized) half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
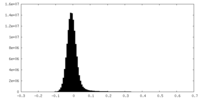
| Density Histograms |
- Sample components
Sample components
-Entire : FljM flagellar filament (symmetrized)
| Entire | Name: FljM flagellar filament (symmetrized) |
|---|---|
| Components |
|
-Supramolecule #1: FljM flagellar filament (symmetrized)
| Supramolecule | Name: FljM flagellar filament (symmetrized) / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Caulobacter vibrioides (bacteria) / Strain: NA1000 Caulobacter vibrioides (bacteria) / Strain: NA1000 |
| Molecular weight | Theoretical: 0.28 kDa/nm |
-Macromolecule #1: Flagellin FljM
| Macromolecule | Name: Flagellin FljM / type: protein_or_peptide / ID: 1 / Number of copies: 44 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Caulobacter vibrioides (bacteria) / Strain: NA1000 Caulobacter vibrioides (bacteria) / Strain: NA1000 |
| Molecular weight | Theoretical: 27.95024 KDa |
| Sequence | String: MALNSINTNS GALIALQNLN STNAELTQVQ QRINTGKKIG SAKDNGAIWA TAKNQSATAG SMNAVKDSLQ RGQSTIDVAL AAGDTITDL LGKMKEKALA ASDTSLNTAS FNALKSDFDS LRDQITKAAS NAKFNGVSIA DGTTTKLSFL ANSDGSAFTV T AKTLTLGG ...String: MALNSINTNS GALIALQNLN STNAELTQVQ QRINTGKKIG SAKDNGAIWA TAKNQSATAG SMNAVKDSLQ RGQSTIDVAL AAGDTITDL LGKMKEKALA ASDTSLNTAS FNALKSDFDS LRDQITKAAS NAKFNGVSIA DGTTTKLSFL ANSDGSAFTV T AKTLTLGG LGLTATSSFT TAAAAKTMIG TIDTALQTAT NKLASLGTSS TGLDTHLTFV GKLQDSLDAG VGNLVDADLA KE SAKLQSL QTKQQLGVQA LSIANQSSSS ILSLFR UniProtKB: Flagellin FljM |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 / Component - Name: phosphate buffered saline |
|---|---|
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 293 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 45.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




 Z
Z Y
Y X
X