+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Membrane protein B | ||||||||||||||||||||||||
 Map data Map data | |||||||||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||||||||
 Keywords Keywords | Membrane Protein / Cryo-EM | ||||||||||||||||||||||||
| Biological species |  Hyphochytrium catenoides (eukaryote) Hyphochytrium catenoides (eukaryote) | ||||||||||||||||||||||||
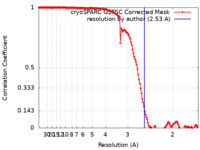
| Method | single particle reconstruction / cryo EM / Resolution: 2.53 Å | ||||||||||||||||||||||||
 Authors Authors | Tajima S / Kim Y / Yamashita K / Fukuda M / Deisseroth K / Kato HE | ||||||||||||||||||||||||
| Funding support |  Japan, 7 items Japan, 7 items
| ||||||||||||||||||||||||
 Citation Citation |  Journal: Cell / Year: 2023 Journal: Cell / Year: 2023Title: Structural basis for ion selectivity in potassium-selective channelrhodopsins. Authors: Seiya Tajima / Yoon Seok Kim / Masahiro Fukuda / YoungJu Jo / Peter Y Wang / Joseph M Paggi / Masatoshi Inoue / Eamon F X Byrne / Koichiro E Kishi / Seiwa Nakamura / Charu Ramakrishnan / ...Authors: Seiya Tajima / Yoon Seok Kim / Masahiro Fukuda / YoungJu Jo / Peter Y Wang / Joseph M Paggi / Masatoshi Inoue / Eamon F X Byrne / Koichiro E Kishi / Seiwa Nakamura / Charu Ramakrishnan / Shunki Takaramoto / Takashi Nagata / Masae Konno / Masahiro Sugiura / Kota Katayama / Toshiki E Matsui / Keitaro Yamashita / Suhyang Kim / Hisako Ikeda / Jaeah Kim / Hideki Kandori / Ron O Dror / Keiichi Inoue / Karl Deisseroth / Hideaki E Kato /    Abstract: KCR channelrhodopsins (K-selective light-gated ion channels) have received attention as potential inhibitory optogenetic tools but more broadly pose a fundamental mystery regarding how their K ...KCR channelrhodopsins (K-selective light-gated ion channels) have received attention as potential inhibitory optogenetic tools but more broadly pose a fundamental mystery regarding how their K selectivity is achieved. Here, we present 2.5-2.7 Å cryo-electron microscopy structures of HcKCR1 and HcKCR2 and of a structure-guided mutant with enhanced K selectivity. Structural, electrophysiological, computational, spectroscopic, and biochemical analyses reveal a distinctive mechanism for K selectivity; rather than forming the symmetrical filter of canonical K channels achieving both selectivity and dehydration, instead, three extracellular-vestibule residues within each monomer form a flexible asymmetric selectivity gate, while a distinct dehydration pathway extends intracellularly. Structural comparisons reveal a retinal-binding pocket that induces retinal rotation (accounting for HcKCR1/HcKCR2 spectral differences), and design of corresponding KCR variants with increased K selectivity (KALI-1/KALI-2) provides key advantages for optogenetic inhibition in vitro and in vivo. Thus, discovery of a mechanism for ion-channel K selectivity also provides a framework for next-generation optogenetics. | ||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34531.map.gz emd_34531.map.gz | 17.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34531-v30.xml emd-34531-v30.xml emd-34531.xml emd-34531.xml | 20.3 KB 20.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_34531_fsc.xml emd_34531_fsc.xml | 13.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_34531.png emd_34531.png | 190.9 KB | ||
| Masks |  emd_34531_msk_1.map emd_34531_msk_1.map | 18.7 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-34531.cif.gz emd-34531.cif.gz | 6.1 KB | ||
| Others |  emd_34531_additional_1.map.gz emd_34531_additional_1.map.gz emd_34531_half_map_1.map.gz emd_34531_half_map_1.map.gz emd_34531_half_map_2.map.gz emd_34531_half_map_2.map.gz | 16.7 MB 17.3 MB 17.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34531 http://ftp.pdbj.org/pub/emdb/structures/EMD-34531 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34531 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34531 | HTTPS FTP |
-Related structure data
| Related structure data |  8h87MC  8h86C  8iu0C M: atomic model generated by this map C: citing same article ( |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_34531.map.gz / Format: CCP4 / Size: 18.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34531.map.gz / Format: CCP4 / Size: 18.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_34531_msk_1.map emd_34531_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
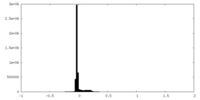
| Density Histograms |
-Additional map: #1
| File | emd_34531_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
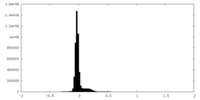
| Density Histograms |
-Half map: #2
| File | emd_34531_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
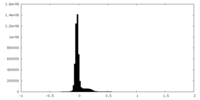
| Density Histograms |
-Half map: #1
| File | emd_34531_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : HcKCR2
| Entire | Name: HcKCR2 |
|---|---|
| Components |
|
-Supramolecule #1: HcKCR2
| Supramolecule | Name: HcKCR2 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Hyphochytrium catenoides (eukaryote) Hyphochytrium catenoides (eukaryote) |
-Macromolecule #1: HcKCR2
| Macromolecule | Name: HcKCR2 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Hyphochytrium catenoides (eukaryote) Hyphochytrium catenoides (eukaryote) |
| Molecular weight | Theoretical: 34.023664 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MPFYDSRPPE GWPRGSVNDM DYPLLGSICA ISAIAIAGSG IWMLYRLDLG MGYSCKPYKS GRAPEVNSIS GIVCLLCGTM YAAKSFDFF DGGGTPFSLN WYWYLDYVFT CPLLIVDFAF TLDIPQKLRY SIAVFVALWC AVAAFATPSA FRFAYYALGC C WFIPLSLF ...String: MPFYDSRPPE GWPRGSVNDM DYPLLGSICA ISAIAIAGSG IWMLYRLDLG MGYSCKPYKS GRAPEVNSIS GIVCLLCGTM YAAKSFDFF DGGGTPFSLN WYWYLDYVFT CPLLIVDFAF TLDIPQKLRY SIAVFVALWC AVAAFATPSA FRFAYYALGC C WFIPLSLF LIRDVKKRYQ VYPPKCQRLL FWACVVFFGF WPLFPLLFIF SWQGSGHISR QAYYIIHAFL DLVCKSIFGF LM TFFRLEL EEHTEVQGLP LKEPKVMDAA AKSRITSEGE YIPLDQIDIN VGAPGSSLEV LFQ |
-Macromolecule #2: RETINAL
| Macromolecule | Name: RETINAL / type: ligand / ID: 2 / Number of copies: 1 / Formula: RET |
|---|---|
| Molecular weight | Theoretical: 284.436 Da |
| Chemical component information |  ChemComp-RET: |
-Macromolecule #3: (7R,17E,20E)-4-HYDROXY-N,N,N-TRIMETHYL-9-OXO-7-[(PALMITOYLOXY)MET...
| Macromolecule | Name: (7R,17E,20E)-4-HYDROXY-N,N,N-TRIMETHYL-9-OXO-7-[(PALMITOYLOXY)METHYL]-3,5,8-TRIOXA-4-PHOSPHAHEXACOSA-17,20-DIEN-1-AMINIUM 4-OXIDE type: ligand / ID: 3 / Number of copies: 6 / Formula: PSC |
|---|---|
| Molecular weight | Theoretical: 759.068 Da |
| Chemical component information |  ChemComp-PSC: |
-Macromolecule #4: PALMITIC ACID
| Macromolecule | Name: PALMITIC ACID / type: ligand / ID: 4 / Number of copies: 9 / Formula: PLM |
|---|---|
| Molecular weight | Theoretical: 256.424 Da |
| Chemical component information |  ChemComp-PLM: |
-Macromolecule #5: water
| Macromolecule | Name: water / type: ligand / ID: 5 / Number of copies: 49 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Sugar embedding | Material: Lipid / Details: nanodisc composing of MSP1D1E3 and soybean lipid |
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 51.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.6 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)