+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of human CEPT1 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CEPT1 / choline/ethanolamine phosphotransferase / LIPID BINDING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationethanolaminephosphotransferase / 1-alkenyl-2-acylglycerol choline phosphotransferase / 1-alkenyl-2-acylglycerol choline phosphotransferase activity / diacylglycerol cholinephosphotransferase / diacylglycerol cholinephosphotransferase activity / ethanolaminephosphotransferase activity / Synthesis of PE / phosphatidylethanolamine biosynthetic process / phosphatidylcholine biosynthetic process / Synthesis of PC ...ethanolaminephosphotransferase / 1-alkenyl-2-acylglycerol choline phosphotransferase / 1-alkenyl-2-acylglycerol choline phosphotransferase activity / diacylglycerol cholinephosphotransferase / diacylglycerol cholinephosphotransferase activity / ethanolaminephosphotransferase activity / Synthesis of PE / phosphatidylethanolamine biosynthetic process / phosphatidylcholine biosynthetic process / Synthesis of PC / lipid metabolic process / nuclear membrane / endoplasmic reticulum membrane / Golgi apparatus / membrane / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.7 Å | |||||||||
 Authors Authors | Qian HW / Wang ZH | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Structural basis for catalysis of human choline/ethanolamine phosphotransferase 1. Authors: Zhenhua Wang / Meng Yang / Yufan Yang / Yonglin He / Hongwu Qian /  Abstract: Phosphatidylcholine (PC) and phosphatidylethanolamine (PE) are two primary components of the eukaryotic membrane and play essential roles in the maintenance of membrane integrity, lipid droplet ...Phosphatidylcholine (PC) and phosphatidylethanolamine (PE) are two primary components of the eukaryotic membrane and play essential roles in the maintenance of membrane integrity, lipid droplet biogenesis, autophagosome formation, and lipoprotein formation and secretion. Choline/ethanolamine phosphotransferase 1 (CEPT1) catalyzes the last step of the biosynthesis of PC and PE in the Kennedy pathway by transferring the substituted phosphate group from CDP-choline/ethanolamine to diacylglycerol. Here, we present the cryo-EM structures of human CEPT1 and its complex with CDP-choline at resolutions of 3.7 Å and 3.8 Å, respectively. CEPT1 is a dimer with 10 transmembrane segments (TMs) in each protomer. TMs 1-6 constitute a conserved catalytic domain with an interior hydrophobic chamber accommodating a PC-like density. Structural observations and biochemical characterizations suggest that the hydrophobic chamber coordinates the acyl tails during the catalytic process. The PC-like density disappears in the structure of the complex with CDP-choline, suggesting a potential substrate-triggered product release mechanism. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34379.map.gz emd_34379.map.gz | 28.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34379-v30.xml emd-34379-v30.xml emd-34379.xml emd-34379.xml | 13.8 KB 13.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_34379.png emd_34379.png | 61.6 KB | ||
| Filedesc metadata |  emd-34379.cif.gz emd-34379.cif.gz | 5.4 KB | ||
| Others |  emd_34379_half_map_1.map.gz emd_34379_half_map_1.map.gz emd_34379_half_map_2.map.gz emd_34379_half_map_2.map.gz | 27.8 MB 27.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34379 http://ftp.pdbj.org/pub/emdb/structures/EMD-34379 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34379 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34379 | HTTPS FTP |
-Validation report
| Summary document |  emd_34379_validation.pdf.gz emd_34379_validation.pdf.gz | 753.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_34379_full_validation.pdf.gz emd_34379_full_validation.pdf.gz | 753.2 KB | Display | |
| Data in XML |  emd_34379_validation.xml.gz emd_34379_validation.xml.gz | 10.8 KB | Display | |
| Data in CIF |  emd_34379_validation.cif.gz emd_34379_validation.cif.gz | 12.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34379 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34379 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34379 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34379 | HTTPS FTP |
-Related structure data
| Related structure data |  8gyxMC  8gywC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_34379.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34379.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||
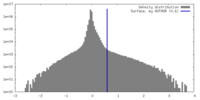
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_34379_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
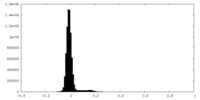
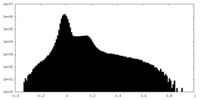
| Density Histograms |
-Half map: #1
| File | emd_34379_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
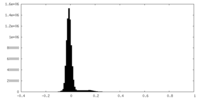
| Density Histograms |
- Sample components
Sample components
-Entire : Choline/ethanolaminephosphotransferase 1 complexed with PC
| Entire | Name: Choline/ethanolaminephosphotransferase 1 complexed with PC |
|---|---|
| Components |
|
-Supramolecule #1: Choline/ethanolaminephosphotransferase 1 complexed with PC
| Supramolecule | Name: Choline/ethanolaminephosphotransferase 1 complexed with PC type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Choline/ethanolaminephosphotransferase 1
| Macromolecule | Name: Choline/ethanolaminephosphotransferase 1 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO / EC number: ethanolaminephosphotransferase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 42.69509 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: TGCVLNKLFQ LPTPPLSRHQ LKRLEEHRYQ SAGRSLLEPL MQGYWEWLVR RVPSWIAPNL ITIIGLSINI CTTILLVFYC PTATEQAPL WAYIACACGL FIYQSLDAID GKQARRTNSS SPLGELFDHG CDSLSTVFVV LGTCIAVQLG TNPDWMFFCC F AGTFMFYC ...String: TGCVLNKLFQ LPTPPLSRHQ LKRLEEHRYQ SAGRSLLEPL MQGYWEWLVR RVPSWIAPNL ITIIGLSINI CTTILLVFYC PTATEQAPL WAYIACACGL FIYQSLDAID GKQARRTNSS SPLGELFDHG CDSLSTVFVV LGTCIAVQLG TNPDWMFFCC F AGTFMFYC AHWQTYVSGT LRFGIIDVTE VQIFIIIMHL LAVIGGPPFW QSMIPVLNIQ MKIFPALCTV AGTIFSCTNY FR VIFTGGV GKNGSTIAGT SVLSPFLHIG SVITLAAMIY KKSAVQLFEK HPCLYILTFG FVSAKITNKL VVAHMTKSEM HLH DTAFIG PALLFLDQYF NSFIDEYIVL WIALVFSFFD LIRYCVSVCN QIASHLHIHV FRIK UniProtKB: Choline/ethanolaminephosphotransferase 1 |
-Macromolecule #2: (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(tri...
| Macromolecule | Name: (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(trimethylammonio)ethyl phosphate type: ligand / ID: 2 / Number of copies: 2 / Formula: POV |
|---|---|
| Molecular weight | Theoretical: 760.076 Da |
| Chemical component information |  ChemComp-POV: |
-Macromolecule #3: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 3 / Number of copies: 4 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 15 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.7 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 851097 |
| Initial angle assignment | Type: ANGULAR RECONSTITUTION |
| Final angle assignment | Type: ANGULAR RECONSTITUTION |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)