[English] 日本語
 Yorodumi
Yorodumi- EMDB-33707: Apoferritin structure at 1.46 angstrom resolution by CryoARM300 e... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Apoferritin structure at 1.46 angstrom resolution by CryoARM300 equipped with Apollo | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 1.47 Å | |||||||||
 Authors Authors | Bammes B / Spilman M / Sakamoto M / Fukumura T / Konyuba Y / Okunishi E | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Apoferritin structure at 1.46 angstrom resolution by CryoARM300 equipped with Apollo Authors: Bammes B / Spilman M / Sakamoto M / Fukumura T / Konyuba Y / Okunishi E | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33707.map.gz emd_33707.map.gz | 17.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33707-v30.xml emd-33707-v30.xml emd-33707.xml emd-33707.xml | 15.3 KB 15.3 KB | Display Display |  EMDB header EMDB header |
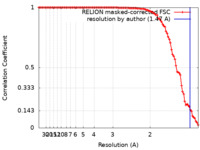
| FSC (resolution estimation) |  emd_33707_fsc.xml emd_33707_fsc.xml | 10.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_33707.png emd_33707.png | 307.9 KB | ||
| Masks |  emd_33707_msk_1.map emd_33707_msk_1.map | 91.1 MB |  Mask map Mask map | |
| Others |  emd_33707_half_map_1.map.gz emd_33707_half_map_1.map.gz emd_33707_half_map_2.map.gz emd_33707_half_map_2.map.gz | 66.8 MB 66.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33707 http://ftp.pdbj.org/pub/emdb/structures/EMD-33707 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33707 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33707 | HTTPS FTP |
-Related structure data
| EM raw data |  EMPIAR-11101 (Title: Apoferritin structure at 1.46 angstrom resolution by CryoARM300II equipped with Apollo EMPIAR-11101 (Title: Apoferritin structure at 1.46 angstrom resolution by CryoARM300II equipped with ApolloData size: 1.1 TB Data #1: Apoferritin structure at 1.46 angstrom resolution by CryoARM300II equipped with Apollo [micrographs - multiframe]) |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_33707.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33707.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.6975 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_33707_msk_1.map emd_33707_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
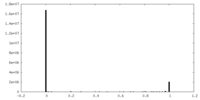
| Density Histograms |
-Half map: #1
| File | emd_33707_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_33707_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Apoferritin
| Entire | Name: Apoferritin |
|---|---|
| Components |
|
-Supramolecule #1: Apoferritin
| Supramolecule | Name: Apoferritin / type: complex / Chimera: Yes / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
-Macromolecule #1: Apoferritin
| Macromolecule | Name: Apoferritin / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Sequence | String: MTTASPSQVR QNYHQDAEAA INRQINLELY ASYVYLSMSC YFDRDDVALK NFAKYFLHQS HEEREHAEKL MKLQNQRGGR IFLQDIKKPD RDDWESGLNA MECALHLEKS VNQSLLELHK LATDKNDPHL CDFIETYYLS EQVKSIKELG DHVTNLRKMG APEAGMAEYL FDKHTLGHGD ES |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 10 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| |||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 80 % / Chamber temperature: 293 K / Instrument: LEICA EM GP |
- Electron microscopy
Electron microscopy
| Microscope | JEOL CRYO ARM 300 |
|---|---|
| Image recording | Film or detector model: OTHER / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 20.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.2 µm / Nominal defocus min: 0.9 µm |
| Sample stage | Specimen holder model: JEOL CRYOSPECPORTER / Cooling holder cryogen: NITROGEN |
 Movie
Movie Controller
Controller



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)