+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Subtomogram average of a yeast nucleosome from BY4741 cells | ||||||||||||||||||||||||||||||
 Map data Map data | Class 1 | ||||||||||||||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||||||||||||||
 Keywords Keywords | Cell / yeast / nucleus / nucleosome / H2A / NUCLEAR PROTEIN / GFP | ||||||||||||||||||||||||||||||
| Biological species |  | ||||||||||||||||||||||||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 24.0 Å | ||||||||||||||||||||||||||||||
 Authors Authors | Tan ZY / Cai S / Noble AJ / Chen JK / Shi J / Gan L | ||||||||||||||||||||||||||||||
| Funding support |  Singapore, Singapore,  United States, 9 items United States, 9 items
| ||||||||||||||||||||||||||||||
 Citation Citation |  Journal: Elife / Year: 2023 Journal: Elife / Year: 2023Title: Heterogeneous non-canonical nucleosomespredominate in yeast cells in situ Authors: Tan ZY / Cai S / Noble AJ / Chen JK / Shi J / Gan L | ||||||||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_31086.map.gz emd_31086.map.gz | 140 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-31086-v30.xml emd-31086-v30.xml emd-31086.xml emd-31086.xml | 22.8 KB 22.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_31086.png emd_31086.png | 32.9 KB | ||
| Others |  emd_31086_additional_1.map.gz emd_31086_additional_1.map.gz emd_31086_additional_2.map.gz emd_31086_additional_2.map.gz emd_31086_additional_3.map.gz emd_31086_additional_3.map.gz emd_31086_half_map_1.map.gz emd_31086_half_map_1.map.gz emd_31086_half_map_2.map.gz emd_31086_half_map_2.map.gz | 141.8 KB 140.9 KB 141.6 KB 141.5 KB 141.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-31086 http://ftp.pdbj.org/pub/emdb/structures/EMD-31086 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31086 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31086 | HTTPS FTP |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_31086.map.gz / Format: CCP4 / Size: 182.6 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_31086.map.gz / Format: CCP4 / Size: 182.6 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Class 1 | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 6.8 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Class 2, half map 1
| File | emd_31086_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Class 2, half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
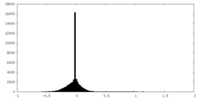
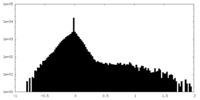
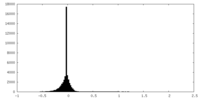
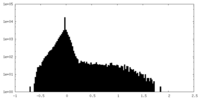
| Density Histograms |
-Additional map: Class 2
| File | emd_31086_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Class 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Class 2, half map 2
| File | emd_31086_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Class 2, half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Class 1, half map 1
| File | emd_31086_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Class 1, half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Class 1, half map 2
| File | emd_31086_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Class 1, half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Nucleosome from budding yeast BY4741
| Entire | Name: Nucleosome from budding yeast BY4741 |
|---|---|
| Components |
|
-Supramolecule #1: Nucleosome from budding yeast BY4741
| Supramolecule | Name: Nucleosome from budding yeast BY4741 / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Grid | Model: C-flat / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY |
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: blot time: 1 second, blot force: 1. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Phase plate: VOLTA PHASE PLATE / Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Detector mode: INTEGRATING / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Number grids imaged: 1 / Average electron dose: 2.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 30369 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 0.1 µm / Nominal defocus min: 0.1 µm / Nominal magnification: 18000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Number classes used: 1 / Applied symmetry - Point group: C1 (asymmetric) / Algorithm: FOURIER SPACE / Resolution.type: BY AUTHOR / Resolution: 24.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: RELION (ver. 3.0.8) / Number subtomograms used: 453 |
|---|---|
| Extraction | Number tomograms: 5 / Number images used: 129473 / Reference model: Featureless cylinder / Method: Template matching / Software - Name: PEET (ver. 1.15) |
| Final 3D classification | Number classes: 2 / Software - Name: RELION (ver. 3.0.8) |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 3.0.8) |
 Movie
Movie Controller
Controller



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)