[English] 日本語
 Yorodumi
Yorodumi- EMDB-23622: Cryo-electron tomogram of FIB-milled Caulobacter crescentus expre... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
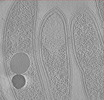
| Title | Cryo-electron tomogram of FIB-milled Caulobacter crescentus expressing WT PopZ | ||||||||||||
 Map data Map data | Caulobacter crescentus cells expressing wildtype PopZ | ||||||||||||
 Sample Sample |
| ||||||||||||
| Biological species |  Caulobacter vibrioides NA1000 (bacteria) Caulobacter vibrioides NA1000 (bacteria) | ||||||||||||
| Method | electron tomography / cryo EM | ||||||||||||
 Authors Authors | Lasker K / Boeynaems S / Lam V / Scholl D / Stainton E / Briner A / Jacquemyn M / Daelemans D / Deniz A / Villa E ...Lasker K / Boeynaems S / Lam V / Scholl D / Stainton E / Briner A / Jacquemyn M / Daelemans D / Deniz A / Villa E / Holehouse AS / Gitler AD / Shapiro L | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: The material properties of a bacterial-derived biomolecular condensate tune biological function in natural and synthetic systems Authors: Lasker K / Boeynaems S / Lam V / Scholl D / Stainton E / Briner A / Jacquemyn M / Daelemans D / Deniz A / Villa E / Holehouse AS / Gitler AD / Shapiro L #1:  Journal: biorxiv / Year: 2021 Journal: biorxiv / Year: 2021Title: A modular platform for engineering function of natural and synthetic biomolecular condensates Authors: Lasker K / Boeynaems S / Lam V / Stainton E / Jacquemyn M / Daelemans D / Villa E / Holehouse AS / Gitler AD / Shapiro L | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_23622.map.gz emd_23622.map.gz | 93.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-23622-v30.xml emd-23622-v30.xml emd-23622.xml emd-23622.xml | 13.2 KB 13.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_23622.png emd_23622.png | 271.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-23622 http://ftp.pdbj.org/pub/emdb/structures/EMD-23622 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23622 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23622 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_23622.map.gz / Format: CCP4 / Size: 152.9 MB / Type: IMAGE STORED AS SIGNED BYTE Download / File: emd_23622.map.gz / Format: CCP4 / Size: 152.9 MB / Type: IMAGE STORED AS SIGNED BYTE | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Caulobacter crescentus cells expressing wildtype PopZ | ||||||||||||||||||||
| Voxel size | X=Y=Z: 17.32 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Caulobacter crescentus cells expressing WT PopZ
| Entire | Name: Caulobacter crescentus cells expressing WT PopZ |
|---|---|
| Components |
|
-Supramolecule #1: Caulobacter crescentus cells expressing WT PopZ
| Supramolecule | Name: Caulobacter crescentus cells expressing WT PopZ / type: cell / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  Caulobacter vibrioides NA1000 (bacteria) Caulobacter vibrioides NA1000 (bacteria) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | electron tomography |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 / Details: M2 minimal media, supplemented with glucose. |
|---|---|
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.019 kPa / Details: 20 mA current. Instrumet: PELCO easiGlow |
| Vitrification | Cryogen name: ETHANE-PROPANE / Instrument: HOMEMADE PLUNGER |
| Details | Protein expression was induced 4-5 hours prior to plunge freezing. Cells were concentrated to an OD600 ~3.0 by centrifugation. |
| Cryo protectant | Trehalose |
| Sectioning | Focused ion beam - Instrument: OTHER / Focused ion beam - Ion: OTHER / Focused ion beam - Voltage: 30 kV / Focused ion beam - Current: 0.01 nA / Focused ion beam - Duration: 2700 sec. / Focused ion beam - Temperature: 77 K / Focused ion beam - Initial thickness: 500 nm / Focused ion beam - Final thickness: 150 nm Focused ion beam - Details: milling current varies from 0.5 nA to 10 pA during milling.. The value given for _emd_sectioning_focused_ion_beam.instrument is FEI Aquilos Cryo-FIB. This is not in a list ...Focused ion beam - Details: milling current varies from 0.5 nA to 10 pA during milling.. The value given for _emd_sectioning_focused_ion_beam.instrument is FEI Aquilos Cryo-FIB. This is not in a list of allowed values {'OTHER', 'DB235'} so OTHER is written into the XML file. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 77.0 K |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 3708 pixel / Digitization - Dimensions - Height: 3838 pixel / Digitization - Frames/image: 1-15 / Number real images: 61 / Average exposure time: 1.45 sec. / Average electron dose: 2.16 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus min: 5.0 µm / Nominal magnification: 33000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Algorithm: BACK PROJECTION / Software - Name:  IMOD (ver. 4.10.29) / Number images used: 61 IMOD (ver. 4.10.29) / Number images used: 61 |
|---|
 Movie
Movie Controller
Controller