[English] 日本語
 Yorodumi
Yorodumi- EMDB-18069: Outward-facing, open1 proteoliposome complex I at 2.8 A, after de... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Outward-facing, open1 proteoliposome complex I at 2.8 A, after deactivation treatment. Initially purified in LMNG. | |||||||||
 Map data Map data | Locally sharpened consensus map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Complex I / Oxidoreductase / Proteoliposomes / Membrane-bound / metabolism / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationComplex I biogenesis / Mitochondrial protein import / RHOG GTPase cycle / Respiratory electron transport / cellular response to oxygen levels / ubiquinone biosynthetic process / mitochondrial large ribosomal subunit binding / gliogenesis / Mitochondrial translation termination / Neutrophil degranulation ...Complex I biogenesis / Mitochondrial protein import / RHOG GTPase cycle / Respiratory electron transport / cellular response to oxygen levels / ubiquinone biosynthetic process / mitochondrial large ribosomal subunit binding / gliogenesis / Mitochondrial translation termination / Neutrophil degranulation / neural precursor cell proliferation / oxygen sensor activity / [2Fe-2S] cluster assembly / Mitochondrial protein degradation / NADH:ubiquinone reductase (H+-translocating) / ubiquinone binding / mitochondrial ATP synthesis coupled electron transport / mitochondrial electron transport, NADH to ubiquinone / electron transport coupled proton transport / acyl binding / mitochondrial respiratory chain complex I assembly / NADH dehydrogenase activity / oxidoreductase activity, acting on NAD(P)H / respiratory chain complex I / NADH dehydrogenase (ubiquinone) activity / acyl carrier activity / response to cAMP / quinone binding / ATP synthesis coupled electron transport / reactive oxygen species metabolic process / neurogenesis / fatty acid binding / aerobic respiration / respiratory electron transport chain / circadian rhythm / brain development / electron transport chain / mitochondrial intermembrane space / 2 iron, 2 sulfur cluster binding / mitochondrial membrane / NAD binding / fatty acid biosynthetic process / FMN binding / 4 iron, 4 sulfur cluster binding / oxidoreductase activity / mitochondrial inner membrane / mitochondrial matrix / negative regulation of DNA-templated transcription / apoptotic process / protein-containing complex binding / mitochondrion / nucleoplasm / metal ion binding / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
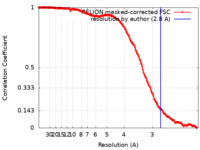
| Method | single particle reconstruction / cryo EM / Resolution: 2.8 Å | |||||||||
 Authors Authors | Grba DN / Hirst J | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2024 Journal: Science / Year: 2024Title: Molecular mechanism of the ischemia-induced regulatory switch in mammalian complex I. Authors: Daniel N Grba / John J Wright / Zhan Yin / William Fisher / Judy Hirst /  Abstract: Respiratory complex I is an efficient driver for oxidative phosphorylation in mammalian mitochondria, but its uncontrolled catalysis under challenging conditions leads to oxidative stress and ...Respiratory complex I is an efficient driver for oxidative phosphorylation in mammalian mitochondria, but its uncontrolled catalysis under challenging conditions leads to oxidative stress and cellular damage. Ischemic conditions switch complex I from rapid, reversible catalysis into a dormant state that protects upon reoxygenation, but the molecular basis for the switch is unknown. We combined precise biochemical definition of complex I catalysis with high-resolution cryo-electron microscopy structures in the phospholipid bilayer of coupled vesicles to reveal the mechanism of the transition into the dormant state, modulated by membrane interactions. By implementing a versatile membrane system to unite structure and function, attributing catalytic and regulatory properties to specific structural states, we define how a conformational switch in complex I controls its physiological roles. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18069.map.gz emd_18069.map.gz | 337 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18069-v30.xml emd-18069-v30.xml emd-18069.xml emd-18069.xml | 82.2 KB 82.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_18069_fsc.xml emd_18069_fsc.xml | 16.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_18069.png emd_18069.png | 101.7 KB | ||
| Masks |  emd_18069_msk_1.map emd_18069_msk_1.map | 421.9 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-18069.cif.gz emd-18069.cif.gz | 16.2 KB | ||
| Others |  emd_18069_additional_1.map.gz emd_18069_additional_1.map.gz emd_18069_additional_2.map.gz emd_18069_additional_2.map.gz emd_18069_half_map_1.map.gz emd_18069_half_map_1.map.gz emd_18069_half_map_2.map.gz emd_18069_half_map_2.map.gz | 24.6 MB 339 MB 340 MB 339.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18069 http://ftp.pdbj.org/pub/emdb/structures/EMD-18069 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18069 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18069 | HTTPS FTP |
-Related structure data
| Related structure data |  8q25MC  8q0aC  8q0fC  8q0jC  8q0mC  8q0oC  8q0qC  8q1pC  8q1uC  8q1yC  8q45C  8q46C  8q47C  8q48C  8q49C  8q4aC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_18069.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_18069.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Locally sharpened consensus map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.072 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_18069_msk_1.map emd_18069_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Globally sharpened/potprocessed consensus map
| File | emd_18069_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Globally sharpened/potprocessed consensus map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Unfiltered consensus map
| File | emd_18069_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unfiltered consensus map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 1
| File | emd_18069_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 2
| File | emd_18069_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Complex I from Bos taurus reconstituted into proteoliposome and t...
+Supramolecule #1: Complex I from Bos taurus reconstituted into proteoliposome and t...
+Macromolecule #1: NADH-ubiquinone oxidoreductase chain 3
+Macromolecule #2: NADH dehydrogenase [ubiquinone] iron-sulfur protein 7, mitochondrial
+Macromolecule #3: NADH dehydrogenase [ubiquinone] iron-sulfur protein 3, mitochondrial
+Macromolecule #4: NADH dehydrogenase [ubiquinone] iron-sulfur protein 2, mitochondrial
+Macromolecule #5: NADH dehydrogenase [ubiquinone] flavoprotein 2, mitochondrial
+Macromolecule #6: NADH dehydrogenase [ubiquinone] flavoprotein 1, mitochondrial
+Macromolecule #7: NADH-ubiquinone oxidoreductase 75 kDa subunit, mitochondrial
+Macromolecule #8: NADH-ubiquinone oxidoreductase chain 1
+Macromolecule #9: NADH dehydrogenase [ubiquinone] iron-sulfur protein 8, mitochondrial
+Macromolecule #10: NADH-ubiquinone oxidoreductase chain 6
+Macromolecule #11: NADH-ubiquinone oxidoreductase chain 4L
+Macromolecule #12: NADH-ubiquinone oxidoreductase chain 5
+Macromolecule #13: NADH-ubiquinone oxidoreductase chain 4
+Macromolecule #14: NADH-ubiquinone oxidoreductase chain 2
+Macromolecule #15: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 10, mi...
+Macromolecule #16: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 9, mit...
+Macromolecule #17: NADH dehydrogenase [ubiquinone] iron-sulfur protein 4, mitochondrial
+Macromolecule #18: NADH dehydrogenase [ubiquinone] iron-sulfur protein 6, mitochondrial
+Macromolecule #19: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 2
+Macromolecule #20: Acyl carrier protein, mitochondrial
+Macromolecule #21: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 5
+Macromolecule #22: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 6
+Macromolecule #23: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 8
+Macromolecule #24: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 11
+Macromolecule #25: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 13
+Macromolecule #26: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 1
+Macromolecule #27: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 3
+Macromolecule #28: NADH dehydrogenase [ubiquinone] 1 subunit C1, mitochondrial
+Macromolecule #29: NADH dehydrogenase [ubiquinone] 1 subunit C2
+Macromolecule #30: NADH dehydrogenase [ubiquinone] iron-sulfur protein 5
+Macromolecule #31: NADH dehydrogenase [ubiquinone] 1 beta subcomplex subunit 1
+Macromolecule #32: NADH dehydrogenase [ubiquinone] 1 beta subcomplex subunit 11, mit...
+Macromolecule #33: NADH dehydrogenase [ubiquinone] 1 beta subcomplex subunit 5, mito...
+Macromolecule #34: NADH dehydrogenase [ubiquinone] 1 beta subcomplex subunit 6
+Macromolecule #35: NADH dehydrogenase [ubiquinone] 1 beta subcomplex subunit 2, mito...
+Macromolecule #36: NADH dehydrogenase [ubiquinone] 1 beta subcomplex subunit 3
+Macromolecule #37: NADH dehydrogenase [ubiquinone] 1 beta subcomplex subunit 8, mito...
+Macromolecule #38: NADH dehydrogenase [ubiquinone] 1 beta subcomplex subunit 4
+Macromolecule #39: NADH dehydrogenase [ubiquinone] 1 beta subcomplex subunit 9
+Macromolecule #40: NADH dehydrogenase [ubiquinone] 1 beta subcomplex subunit 7
+Macromolecule #41: NADH dehydrogenase [ubiquinone] 1 beta subcomplex subunit 10
+Macromolecule #42: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 12
+Macromolecule #43: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 7
+Macromolecule #44: NADH dehydrogenase [ubiquinone] flavoprotein 3, mitochondrial
+Macromolecule #45: 1,2-Distearoyl-sn-glycerophosphoethanolamine
+Macromolecule #46: 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE
+Macromolecule #47: IRON/SULFUR CLUSTER
+Macromolecule #48: FE2/S2 (INORGANIC) CLUSTER
+Macromolecule #49: FLAVIN MONONUCLEOTIDE
+Macromolecule #50: POTASSIUM ION
+Macromolecule #51: UBIQUINONE-10
+Macromolecule #52: CARDIOLIPIN
+Macromolecule #53: DIUNDECYL PHOSPHATIDYL CHOLINE
+Macromolecule #54: ZINC ION
+Macromolecule #55: MAGNESIUM ION
+Macromolecule #56: 2'-DEOXYGUANOSINE-5'-TRIPHOSPHATE
+Macromolecule #57: NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE
+Macromolecule #58: ~{S}-[2-[3-[[(2~{R})-3,3-dimethyl-2-oxidanyl-4-phosphonooxy-butan...
+Macromolecule #59: CHOLIC ACID
+Macromolecule #60: MYRISTIC ACID
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.2 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| |||||||||
| Grid | Model: UltrAuFoil / Material: GOLD / Support film - Material: GRAPHENE OXIDE / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 90 sec. / Pretreatment - Atmosphere: AIR | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK III Details: Sample was incubated on grid for 30 s before plunge freezing. | |||||||||
| Details | Complex I proteoliposomes on a graphene-oxide coated gold grid with even liposome distribution. Sample was deactivated with heat treatment prior to freezing. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | #0 - Image recording ID: 1 / #0 - Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / #0 - Digitization - Dimensions - Width: 5760 pixel / #0 - Digitization - Dimensions - Height: 4092 pixel / #0 - Number grids imaged: 1 / #0 - Number real images: 6692 / #0 - Average exposure time: 3.8 sec. / #0 - Average electron dose: 40.0 e/Å2 #0 - Details: The data were collected with aberration-free image shift (AFIS). #1 - Image recording ID: 2 / #1 - Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / #1 - Digitization - Dimensions - Width: 5760 pixel / #1 - Digitization - Dimensions - Height: 4092 pixel / #1 - Number grids imaged: 1 / #1 - Number real images: 4550 / #1 - Average exposure time: 3.8 sec. / #1 - Average electron dose: 40.0 e/Å2 #1 - Details: The data were collected without aberration-free image shift and with a 20 degree tilt. The same grid as the non-tilted collection was used. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.3000000000000003 µm / Nominal defocus min: 0.9 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)