[English] 日本語
 Yorodumi
Yorodumi- EMDB-17191: The H/ACA RNP lobe of human telomerase with the dyskerin thumb lo... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | The H/ACA RNP lobe of human telomerase with the dyskerin thumb loop in an open conformation | ||||||||||||
 Map data Map data | Final post-processed sharpened map. | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | telomerase / H/ACA / pseudouridylation / ribonucleoprotein / RNA BINDING PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationtelomere formation via telomerase / box H/ACA scaRNP complex / box H/ACA telomerase RNP complex / protein localization to Cajal body / snoRNA guided rRNA pseudouridine synthesis / enzyme-directed rRNA pseudouridine synthesis / pseudouridine synthesis / Cajal body organization / Isomerases; Intramolecular transferases; Transferring other groups / rRNA pseudouridine synthesis ...telomere formation via telomerase / box H/ACA scaRNP complex / box H/ACA telomerase RNP complex / protein localization to Cajal body / snoRNA guided rRNA pseudouridine synthesis / enzyme-directed rRNA pseudouridine synthesis / pseudouridine synthesis / Cajal body organization / Isomerases; Intramolecular transferases; Transferring other groups / rRNA pseudouridine synthesis / box H/ACA snoRNP complex / box H/ACA sno(s)RNA 3'-end processing / telomerase RNA stabilization / protein carrier activity / regulation of telomerase RNA localization to Cajal body / snRNA pseudouridine synthesis / mRNA pseudouridine synthesis / box H/ACA snoRNA binding / pseudouridine synthase activity / telomerase RNA localization to Cajal body / telomerase activity / positive regulation of establishment of protein localization to telomere / scaRNA localization to Cajal body / positive regulation of telomerase RNA localization to Cajal body / RNA folding chaperone / sno(s)RNA-containing ribonucleoprotein complex / telomerase holoenzyme complex / telomerase RNA binding / U3 snoRNA binding / rRNA modification in the nucleus and cytosol / positive regulation of double-strand break repair via nonhomologous end joining / telomerase holoenzyme complex assembly / positive regulation of double-strand break repair / Association of TriC/CCT with target proteins during biosynthesis / Telomere Extension By Telomerase / RNA folding / telomere maintenance via telomerase / Cajal body / RNA processing / positive regulation of double-strand break repair via homologous recombination / positive regulation of telomere maintenance via telomerase / positive regulation of DNA repair / mRNA 3'-UTR binding / fibrillar center / rRNA processing / site of double-strand break / protein-folding chaperone binding / histone binding / chromosome, telomeric region / nuclear body / DNA repair / ubiquitin protein ligase binding / protein-containing complex binding / nucleolus / RNA binding / nucleoplasm / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
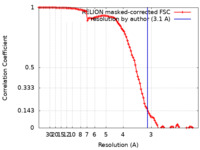
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | ||||||||||||
 Authors Authors | Ghanim GE / Sekne Z / van Roon AMM / Balch S / Nguyen THD | ||||||||||||
| Funding support |  United Kingdom, United Kingdom,  United States, European Union, 3 items United States, European Union, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: 2.7 Å cryo-EM structure of human telomerase H/ACA ribonucleoprotein. Authors: George E Ghanim / Zala Sekne / Sebastian Balch / Anne-Marie M van Roon / Thi Hoang Duong Nguyen /  Abstract: Telomerase is a ribonucleoprotein (RNP) enzyme that extends telomeric repeats at eukaryotic chromosome ends to counterbalance telomere loss caused by incomplete genome replication. Human telomerase ...Telomerase is a ribonucleoprotein (RNP) enzyme that extends telomeric repeats at eukaryotic chromosome ends to counterbalance telomere loss caused by incomplete genome replication. Human telomerase is comprised of two distinct functional lobes tethered by telomerase RNA (hTR): a catalytic core, responsible for DNA extension; and a Hinge and ACA (H/ACA) box RNP, responsible for telomerase biogenesis. H/ACA RNPs also have a general role in pseudouridylation of spliceosomal and ribosomal RNAs, which is critical for the biogenesis of the spliceosome and ribosome. Much of our structural understanding of eukaryotic H/ACA RNPs comes from structures of the human telomerase H/ACA RNP. Here we report a 2.7 Å cryo-electron microscopy structure of the telomerase H/ACA RNP. The significant improvement in resolution over previous 3.3 Å to 8.2 Å structures allows us to uncover new molecular interactions within the H/ACA RNP. Many disease mutations are mapped to these interaction sites. The structure also reveals unprecedented insights into a region critical for pseudouridylation in canonical H/ACA RNPs. Together, our work advances understanding of telomerase-related disease mutations and the mechanism of pseudouridylation by eukaryotic H/ACA RNPs. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17191.map.gz emd_17191.map.gz | 77.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17191-v30.xml emd-17191-v30.xml emd-17191.xml emd-17191.xml | 29.6 KB 29.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_17191_fsc.xml emd_17191_fsc.xml | 10 KB | Display |  FSC data file FSC data file |
| Images |  emd_17191.png emd_17191.png | 67.6 KB | ||
| Filedesc metadata |  emd-17191.cif.gz emd-17191.cif.gz | 8.3 KB | ||
| Others |  emd_17191_additional_1.map.gz emd_17191_additional_1.map.gz emd_17191_half_map_1.map.gz emd_17191_half_map_1.map.gz emd_17191_half_map_2.map.gz emd_17191_half_map_2.map.gz | 65.4 MB 65.7 MB 65.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17191 http://ftp.pdbj.org/pub/emdb/structures/EMD-17191 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17191 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17191 | HTTPS FTP |
-Related structure data
| Related structure data |  8oufMC  8oueC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_17191.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17191.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Final post-processed sharpened map. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.059 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Final unsharped map.
| File | emd_17191_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Final unsharped map. | ||||||||||||
| Projections & Slices |
| ||||||||||||
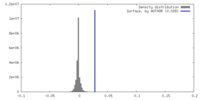
| Density Histograms |
-Half map: Half-map2 resulting from gold standard refinement in RELION.
| File | emd_17191_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map2 resulting from gold standard refinement in RELION. | ||||||||||||
| Projections & Slices |
| ||||||||||||
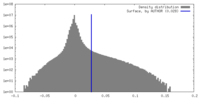
| Density Histograms |
-Half map: Half-map1 resulting from gold standard refinement in RELION.
| File | emd_17191_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map1 resulting from gold standard refinement in RELION. | ||||||||||||
| Projections & Slices |
| ||||||||||||
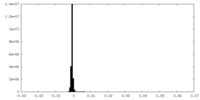
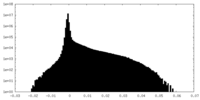
| Density Histograms |
- Sample components
Sample components
-Entire : Human telomerase H/ACA RNP lobe
| Entire | Name: Human telomerase H/ACA RNP lobe |
|---|---|
| Components |
|
-Supramolecule #1: Human telomerase H/ACA RNP lobe
| Supramolecule | Name: Human telomerase H/ACA RNP lobe / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: The hTR H/ACA motif, which adopts a double-hairpin structure hinged by the H-box and flanked by the ACA box, scaffolds the assembly of two copies each of dyskerin, NHP2, NOP10 and GAR1, one ...Details: The hTR H/ACA motif, which adopts a double-hairpin structure hinged by the H-box and flanked by the ACA box, scaffolds the assembly of two copies each of dyskerin, NHP2, NOP10 and GAR1, one on each hairpin. TCAB1 binds the CAB box. |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 415 KDa |
-Supramolecule #2: Telomerase H/ACA RNP lobe
| Supramolecule | Name: Telomerase H/ACA RNP lobe / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #2-#6 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: Human telomerase RNA
| Supramolecule | Name: Human telomerase RNA / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Human telomerase RNA
| Macromolecule | Name: Human telomerase RNA / type: rna / ID: 1 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Organ: Kidney Homo sapiens (human) / Organ: Kidney |
| Molecular weight | Theoretical: 145.477797 KDa |
| Sequence | String: GGGUUGCGGA GGGUGGGCCU GGGAGGGGUG GUGGCCAUUU UUUGUCUAAC CCUAACUGAG AAGGGCGUAG GCGCCGUGCU UUUGCUCCC CGCGCGCUGU UUUUCUCGCU GACUUUCAGC GGGCGGAAAA GCCUCGGCCU GCCGCCUUCC ACCGUUCAUU C UAGAGCAA ...String: GGGUUGCGGA GGGUGGGCCU GGGAGGGGUG GUGGCCAUUU UUUGUCUAAC CCUAACUGAG AAGGGCGUAG GCGCCGUGCU UUUGCUCCC CGCGCGCUGU UUUUCUCGCU GACUUUCAGC GGGCGGAAAA GCCUCGGCCU GCCGCCUUCC ACCGUUCAUU C UAGAGCAA ACAAAAAAUG UCAGCUGCUG GCCCGUUCGC CCCUCCCGGG GACCUGCGGC GGGUCGCCUG CCCAGCCCCC GA ACCCCGC CUGGAGGCCG CGGUCGGCCC GGGGCUUCUC CGGAGGCACC CACUGCCACC GCGAAGAGUU GGGCUCUGUC AGC CGCGGG UCUCUCGGGG GCGAGGGCGA GGUUCAGGCC UUUCAGGCCG CAGGAAGAGG AACGGAGCGA GUCCCCGCGC GCGG CGCGA UUCCCUGAGC UGUGGGACGU GCACCCAGGA CUCGGCUCAC ACAUGC GENBANK: GENBANK: U85256.1 |
-Macromolecule #2: H/ACA ribonucleoprotein complex subunit DKC1
| Macromolecule | Name: H/ACA ribonucleoprotein complex subunit DKC1 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Organ: Kidney Homo sapiens (human) / Organ: Kidney |
| Molecular weight | Theoretical: 57.779211 KDa |
| Sequence | String: MADAEVIILP KKHKKKKERK SLPEEDVAEI QHAEEFLIKP ESKVAKLDTS QWPLLLKNFD KLNVRTTHYT PLACGSNPLK REIGDYIRT GFINLDKPSN PSSHEVVAWI RRILRVEKTG HSGTLDPKVT GCLIVCIERA TRLVKSQQSA GKEYVGIVRL H NAIEGGTQ ...String: MADAEVIILP KKHKKKKERK SLPEEDVAEI QHAEEFLIKP ESKVAKLDTS QWPLLLKNFD KLNVRTTHYT PLACGSNPLK REIGDYIRT GFINLDKPSN PSSHEVVAWI RRILRVEKTG HSGTLDPKVT GCLIVCIERA TRLVKSQQSA GKEYVGIVRL H NAIEGGTQ LSRALETLTG ALFQRPPLIA AVKRQLRVRT IYESKMIEYD PERRLGIFWV SCEAGTYIRT LCVHLGLLLG VG GQMQELR RVRSGVMSEK DHMVTMHDVL DAQWLYDNHK DESYLRRVVY PLEKLLTSHK RLVMKDSAVN AICYGAKIML PGV LRYEDG IEVNQEIVVI TTKGEAICMA IALMTTAVIS TCDHGIVAKI KRVIMERDTY PRKWGLGPKA SQKKLMIKQG LLDK HGKPT DSTPATWKQE YVDYSESAKK EVVAEVVKAP QVVAEAAKTA KRKRESESES DETPPAAPQL IKKEKKKSKK DKKAK AGLE SGAEPGDGDS DTTKKKKKKK KAKEVELVSE UniProtKB: H/ACA ribonucleoprotein complex subunit DKC1 |
-Macromolecule #3: H/ACA ribonucleoprotein complex subunit 1
| Macromolecule | Name: H/ACA ribonucleoprotein complex subunit 1 / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Organ: Kidney Homo sapiens (human) / Organ: Kidney |
| Molecular weight | Theoretical: 22.387963 KDa |
| Sequence | String: MSFRGGGRGG FNRGGGGGGF NRGGSSNHFR GGGGGGGGGN FRGGGRGGFG RGGGRGGFNK GQDQGPPERV VLLGEFLHPC EDDIVCKCT TDENKVPYFN APVYLENKEQ IGKVDEIFGQ LRDFYFSVKL SENMKASSFK KLQKFYIDPY KLLPLQRFLP R PPGEKGPP ...String: MSFRGGGRGG FNRGGGGGGF NRGGSSNHFR GGGGGGGGGN FRGGGRGGFG RGGGRGGFNK GQDQGPPERV VLLGEFLHPC EDDIVCKCT TDENKVPYFN APVYLENKEQ IGKVDEIFGQ LRDFYFSVKL SENMKASSFK KLQKFYIDPY KLLPLQRFLP R PPGEKGPP RGGGRGGRGG GRGGGGRGGG RGGGFRGGRG GGGGGFRGGR GGGFRGRGH UniProtKB: H/ACA ribonucleoprotein complex subunit 1 |
-Macromolecule #4: H/ACA ribonucleoprotein complex subunit 2
| Macromolecule | Name: H/ACA ribonucleoprotein complex subunit 2 / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Organ: Kidney Homo sapiens (human) / Organ: Kidney |
| Molecular weight | Theoretical: 17.22607 KDa |
| Sequence | String: MTKIKADPDG PEAQAEACSG ERTYQELLVN QNPIAQPLAS RRLTRKLYKC IKKAVKQKQI RRGVKEVQKF VNKGEKGIMV LAGDTLPIE VYCHLPVMCE DRNLPYVYIP SKTDLGAAAG SKRPTCVIMV KPHEEYQEAY DECLEEVQSL PLPL UniProtKB: H/ACA ribonucleoprotein complex subunit 2 |
-Macromolecule #5: H/ACA ribonucleoprotein complex subunit 3
| Macromolecule | Name: H/ACA ribonucleoprotein complex subunit 3 / type: protein_or_peptide / ID: 5 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Organ: Kidney Homo sapiens (human) / Organ: Kidney |
| Molecular weight | Theoretical: 7.719989 KDa |
| Sequence | String: MFLQYYLNEQ GDRVYTLKKF DPMGQQTCSA HPARFSPDDK YSRHRITIKK RFKVLMTQQP RPVL UniProtKB: H/ACA ribonucleoprotein complex subunit 3 |
-Macromolecule #6: Telomerase Cajal body protein 1
| Macromolecule | Name: Telomerase Cajal body protein 1 / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Organ: Kidney Homo sapiens (human) / Organ: Kidney |
| Molecular weight | Theoretical: 59.35707 KDa |
| Sequence | String: MKTLETQPLA PDCCPSDQDP APAHPSPHAS PMNKNADSEL MPPPPERGDP PRLSPDPVAG SAVSQELREG DPVSLSTPLE TEFGSPSEL SPRIEEQELS ENTSLPAEEA NGSLSEEEAN GPELGSGKAM EDTSGEPAAE DEGDTAWNYS FSQLPRFLSG S WSEFSTQP ...String: MKTLETQPLA PDCCPSDQDP APAHPSPHAS PMNKNADSEL MPPPPERGDP PRLSPDPVAG SAVSQELREG DPVSLSTPLE TEFGSPSEL SPRIEEQELS ENTSLPAEEA NGSLSEEEAN GPELGSGKAM EDTSGEPAAE DEGDTAWNYS FSQLPRFLSG S WSEFSTQP ENFLKGCKWA PDGSCILTNS ADNILRIYNL PPELYHEGEQ VEYAEMVPVL RMVEGDTIYD YCWYSLMSSA QP DTSYVAS SSRENPIHIW DAFTGELRAS FRAYNHLDEL TAAHSLCFSP DGSQLFCGFN RTVRVFSTAR PGRDCEVRAT FAK KQGQSG IISCIAFSPA QPLYACGSYG RSLGLYAWDD GSPLALLGGH QGGITHLCFH PDGNRFFSGA RKDAELLCWD LRQS GYPLW SLGREVTTNQ RIYFDLDPTG QFLVSGSTSG AVSVWDTDGP GNDGKPEPVL SFLPQKDCTN GVSLHPSLPL LATAS GQRV FPEPTESGDE GEELGLPLLS TRHVHLECRL QLWWCGGAPD SSIPDDHQGE KGQGGTEGGV GELI UniProtKB: Telomerase Cajal body protein 1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 Component:
| |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: C-flat / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 5 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 12 sec. / Pretreatment - Atmosphere: AIR | |||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 78.0 K |
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Number grids imaged: 2 / Number real images: 41053 / Average exposure time: 3.0 sec. / Average electron dose: 48.0 e/Å2 Details: Images were collected in movie-mode and fractionated into 48 movie frames |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated magnification: 45872 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)