[English] 日本語
 Yorodumi
Yorodumi- EMDB-15799: cryo-EM structure of carboxysomal mini-shell: icosahedral assembl... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | cryo-EM structure of carboxysomal mini-shell: icosahedral assembly from CsoS4A/1A and CsoS2 co-expression (T = 4) | |||||||||
 Map data Map data | merged map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | carboxysome / shell / icosahedral symmetry / STRUCTURAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationstructural constituent of carboxysome shell / carboxysome / carbon fixation Similarity search - Function | |||||||||
| Biological species |  Halothiobacillus neapolitanus (bacteria) Halothiobacillus neapolitanus (bacteria) | |||||||||
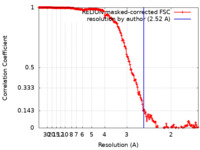
| Method | single particle reconstruction / cryo EM / Resolution: 2.52 Å | |||||||||
 Authors Authors | Ni T / Jiang Q / Liu LN / Zhang P | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Intrinsically disordered CsoS2 acts as a general molecular thread for α-carboxysome shell assembly. Authors: Tao Ni / Qiuyao Jiang / Pei Cing Ng / Juan Shen / Hao Dou / Yanan Zhu / Julika Radecke / Gregory F Dykes / Fang Huang / Lu-Ning Liu / Peijun Zhang /   Abstract: Carboxysomes are a paradigm of self-assembling proteinaceous organelles found in nature, offering compartmentalisation of enzymes and pathways to enhance carbon fixation. In α-carboxysomes, the ...Carboxysomes are a paradigm of self-assembling proteinaceous organelles found in nature, offering compartmentalisation of enzymes and pathways to enhance carbon fixation. In α-carboxysomes, the disordered linker protein CsoS2 plays an essential role in carboxysome assembly and Rubisco encapsulation. Its mechanism of action, however, is not fully understood. Here we synthetically engineer α-carboxysome shells using minimal shell components and determine cryoEM structures of these to decipher the principle of shell assembly and encapsulation. The structures reveal that the intrinsically disordered CsoS2 C-terminus is well-structured and acts as a universal "molecular thread" stitching through multiple shell protein interfaces. We further uncover in CsoS2 a highly conserved repetitive key interaction motif, [IV]TG, which is critical to the shell assembly and architecture. Our study provides a general mechanism for the CsoS2-governed carboxysome shell assembly and cargo encapsulation and further advances synthetic engineering of carboxysomes for diverse biotechnological applications. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15799.map.gz emd_15799.map.gz | 479.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15799-v30.xml emd-15799-v30.xml emd-15799.xml emd-15799.xml | 16.4 KB 16.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15799_fsc.xml emd_15799_fsc.xml | 18 KB | Display |  FSC data file FSC data file |
| Images |  emd_15799.png emd_15799.png | 175.3 KB | ||
| Masks |  emd_15799_msk_1.map emd_15799_msk_1.map | 512 MB |  Mask map Mask map | |
| Others |  emd_15799_half_map_1.map.gz emd_15799_half_map_1.map.gz emd_15799_half_map_2.map.gz emd_15799_half_map_2.map.gz | 411.2 MB 411.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15799 http://ftp.pdbj.org/pub/emdb/structures/EMD-15799 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15799 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15799 | HTTPS FTP |
-Related structure data
| Related structure data |  8b11MC  8b0yC  8b12C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_15799.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15799.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | merged map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.831 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_15799_msk_1.map emd_15799_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
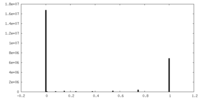
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half1
| File | emd_15799_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half2
| File | emd_15799_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : mini-shell assembly from Halothiobacillus neapolitanus with CsoS4...
| Entire | Name: mini-shell assembly from Halothiobacillus neapolitanus with CsoS4A/1A and CsoS2 co-expression (T= 4) |
|---|---|
| Components |
|
-Supramolecule #1: mini-shell assembly from Halothiobacillus neapolitanus with CsoS4...
| Supramolecule | Name: mini-shell assembly from Halothiobacillus neapolitanus with CsoS4A/1A and CsoS2 co-expression (T= 4) type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Halothiobacillus neapolitanus (bacteria) Halothiobacillus neapolitanus (bacteria) |
-Macromolecule #1: Carboxysome shell vertex protein CsoS4A
| Macromolecule | Name: Carboxysome shell vertex protein CsoS4A / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Halothiobacillus neapolitanus (bacteria) / Strain: ATCC 23641 / c2 Halothiobacillus neapolitanus (bacteria) / Strain: ATCC 23641 / c2 |
| Molecular weight | Theoretical: 8.900287 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKIMQVEKTL VSTNRIADMG HKPLLVVWEK PGAPRQVAVD AIGCIPGDWV LCVGSSAARE AAGSKSYPSD LTIIGIIDQW NGE UniProtKB: Carboxysome shell vertex protein CsoS4A |
-Macromolecule #2: Major carboxysome shell protein CsoS1A
| Macromolecule | Name: Major carboxysome shell protein CsoS1A / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Halothiobacillus neapolitanus (bacteria) / Strain: ATCC 23641 / c2 Halothiobacillus neapolitanus (bacteria) / Strain: ATCC 23641 / c2 |
| Molecular weight | Theoretical: 9.973478 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MADVTGIALG MIETRGLVPA IEAADAMTKA AEVRLVGRQF VGGGYVTVLV RGETGAVNAA VRAGADACER VGDGLVAAHI IARVHSEVE NILPKAPQA UniProtKB: Major carboxysome shell protein CsoS1A |
-Macromolecule #3: water
| Macromolecule | Name: water / type: ligand / ID: 3 / Number of copies: 45 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 44.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)