+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | beta-2-microglobulin DeltaN6 amyloid fibril form 2PFb | ||||||||||||
 Map data Map data | CryoEM helical symmetrised map of b2m-DN6 Form 2PFb (sharpening b-factor of -37) | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Amyloid / fibril / helical / cross-beta / dialysis-related amyloidosis / b2m / polymorph / PROTEIN FIBRIL | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationearly endosome lumen / Nef mediated downregulation of MHC class I complex cell surface expression / DAP12 interactions / Endosomal/Vacuolar pathway / T cell mediated cytotoxicity / Antigen Presentation: Folding, assembly and peptide loading of class I MHC / regulation of iron ion transport / cellular response to iron(III) ion / negative regulation of iron ion transport / negative regulation of forebrain neuron differentiation ...early endosome lumen / Nef mediated downregulation of MHC class I complex cell surface expression / DAP12 interactions / Endosomal/Vacuolar pathway / T cell mediated cytotoxicity / Antigen Presentation: Folding, assembly and peptide loading of class I MHC / regulation of iron ion transport / cellular response to iron(III) ion / negative regulation of iron ion transport / negative regulation of forebrain neuron differentiation / antigen processing and presentation of exogenous protein antigen via MHC class Ib, TAP-dependent / peptide antigen assembly with MHC class I protein complex / ER to Golgi transport vesicle membrane / regulation of erythrocyte differentiation / response to molecule of bacterial origin / HFE-transferrin receptor complex / transferrin transport / MHC class I peptide loading complex / cellular response to iron ion / negative regulation of receptor-mediated endocytosis / positive regulation of T cell cytokine production / antigen processing and presentation of endogenous peptide antigen via MHC class I / MHC class I protein complex / peptide antigen assembly with MHC class II protein complex / negative regulation of neurogenesis / cellular response to nicotine / MHC class II protein complex / positive regulation of receptor-mediated endocytosis / multicellular organismal-level iron ion homeostasis / positive regulation of T cell mediated cytotoxicity / specific granule lumen / antigen processing and presentation of exogenous peptide antigen via MHC class II / positive regulation of immune response / peptide antigen binding / phagocytic vesicle membrane / recycling endosome membrane / positive regulation of T cell activation / negative regulation of epithelial cell proliferation / Interferon gamma signaling / Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell / Modulation by Mtb of host immune system / positive regulation of cellular senescence / sensory perception of smell / tertiary granule lumen / MHC class II protein complex binding / DAP12 signaling / T cell differentiation in thymus / late endosome membrane / negative regulation of neuron projection development / protein refolding / ER-Phagosome pathway / early endosome membrane / amyloid fibril formation / protein homotetramerization / intracellular iron ion homeostasis / learning or memory / endoplasmic reticulum lumen / Amyloid fiber formation / Golgi membrane / external side of plasma membrane / lysosomal membrane / focal adhesion / Neutrophil degranulation / SARS-CoV-2 activates/modulates innate and adaptive immune responses / structural molecule activity / endoplasmic reticulum / Golgi apparatus / protein homodimerization activity / : / extracellular exosome / extracellular region / membrane / identical protein binding / plasma membrane / cytosol Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
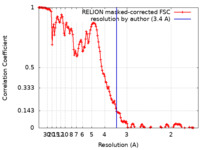
| Method | helical reconstruction / cryo EM / Resolution: 3.4 Å | ||||||||||||
 Authors Authors | Wilkinson M / Gallardo R / Radford SE / Ranson NA | ||||||||||||
| Funding support |  United Kingdom, 3 items United Kingdom, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Disease-relevant β-microglobulin variants share a common amyloid fold. Authors: Martin Wilkinson / Rodrigo U Gallardo / Roberto Maya Martinez / Nicolas Guthertz / Masatomo So / Liam D Aubrey / Sheena E Radford / Neil A Ranson /    Abstract: β-microglobulin (βm) and its truncated variant ΔΝ6 are co-deposited in amyloid fibrils in the joints, causing the disorder dialysis-related amyloidosis (DRA). Point mutations of βm result in ...β-microglobulin (βm) and its truncated variant ΔΝ6 are co-deposited in amyloid fibrils in the joints, causing the disorder dialysis-related amyloidosis (DRA). Point mutations of βm result in diseases with distinct pathologies. βm-D76N causes a rare systemic amyloidosis with protein deposited in the viscera in the absence of renal failure, whilst βm-V27M is associated with renal failure, with amyloid deposits forming predominantly in the tongue. Here we use cryoEM to determine the structures of fibrils formed from these variants under identical conditions in vitro. We show that each fibril sample is polymorphic, with diversity arising from a 'lego-like' assembly of a common amyloid building block. These results suggest a 'many sequences, one amyloid fold' paradigm in contrast with the recently reported 'one sequence, many amyloid folds' behaviour of intrinsically disordered proteins such as tau and Aβ. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15223.map.gz emd_15223.map.gz | 48.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15223-v30.xml emd-15223-v30.xml emd-15223.xml emd-15223.xml | 19.1 KB 19.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15223_fsc.xml emd_15223_fsc.xml | 14.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_15223.png emd_15223.png | 63.1 KB | ||
| Filedesc metadata |  emd-15223.cif.gz emd-15223.cif.gz | 6.3 KB | ||
| Others |  emd_15223_half_map_1.map.gz emd_15223_half_map_1.map.gz emd_15223_half_map_2.map.gz emd_15223_half_map_2.map.gz | 228.5 MB 228.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15223 http://ftp.pdbj.org/pub/emdb/structures/EMD-15223 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15223 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15223 | HTTPS FTP |
-Related structure data
| Related structure data |  8a7pMC  8a7oC  8a7qC  8a7tC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_15223.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15223.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CryoEM helical symmetrised map of b2m-DN6 Form 2PFb (sharpening b-factor of -37) | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: halfmap2
| File | emd_15223_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | halfmap2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: halfmap1
| File | emd_15223_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | halfmap1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Amyloid fibril polymorph 2PFb of the beta-2-microglobulin deltaN6...
| Entire | Name: Amyloid fibril polymorph 2PFb of the beta-2-microglobulin deltaN6 variant. |
|---|---|
| Components |
|
-Supramolecule #1: Amyloid fibril polymorph 2PFb of the beta-2-microglobulin deltaN6...
| Supramolecule | Name: Amyloid fibril polymorph 2PFb of the beta-2-microglobulin deltaN6 variant. type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: Recombinantly expressed and fibrillated in vitro at pH 6.2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Beta-2-microglobulin form pI 5.3
| Macromolecule | Name: Beta-2-microglobulin form pI 5.3 / type: protein_or_peptide / ID: 1 Details: Natural variant deltaN6 with additional methionine added prior to peptide sequence for bacterial expression Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 11.153478 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MIQVYSRHPA ENGKSNFLNC YVSGFHPSDI EVDLLKNGER IEKVEHSDLS FSKDWSFYLL YYTEFTPTEK DEYACRVNHV TLSQPKIVK WDRDM UniProtKB: Beta-2-microglobulin |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 6.2 Component:
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: EMS Lacey Carbon / Material: COPPER / Mesh: 300 / Support film - Material: GRAPHENE OXIDE / Support film - topology: CONTINUOUS / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 60 sec. Details: The grid was plasma cleaned prior to 2x application of graphene oxide-DDM mixture, then grid was used immediately for sample application and vitrification | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: 6s blot. | |||||||||
| Details | Fibrillation conditions: 20 uM monomeric b2m-DN6 at 37C with shaking at 600 rpm for 2-3 weeks |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Number grids imaged: 1 / Number real images: 4095 / Average exposure time: 7.0 sec. / Average electron dose: 43.0 e/Å2 Details: 1687 raw EER frames were collected per image and combined into 40 fractions for processing |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.3 µm / Nominal magnification: 96000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL / Overall B value: 81 |
|---|---|
| Output model |  PDB-8a7p: |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)