[English] 日本語
 Yorodumi
Yorodumi- EMDB-15214: Endogenous yeast L-A helper virus identified from native cell extracts -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Endogenous yeast L-A helper virus identified from native cell extracts | ||||||||||||||||||
 Map data Map data | Icosahedrally averaged capsid of the L-A helper virus from native Saccharomyces cerevisiae cell extract | ||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | Capsid structure ScVLA / viral particle / wildtype / endogenous / VIRUS | ||||||||||||||||||
| Biological species |  Saccharomyces cerevisiae virus L-A Saccharomyces cerevisiae virus L-A | ||||||||||||||||||
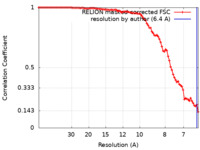
| Method | single particle reconstruction / cryo EM / Resolution: 6.4 Å | ||||||||||||||||||
 Authors Authors | Schmidt L / Kyrilis F / Hamdi F / Semchonok DA / Kastritis PL | ||||||||||||||||||
| Funding support |  Germany, European Union, 5 items Germany, European Union, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Commun Biol / Year: 2024 Journal: Commun Biol / Year: 2024Title: Delineating organizational principles of the endogenous L-A virus by cryo-EM and computational analysis of native cell extracts. Authors: Lisa Schmidt / Christian Tüting / Fotis L Kyrilis / Farzad Hamdi / Dmitry A Semchonok / Gerd Hause / Annette Meister / Christian Ihling / Milton T Stubbs / Andrea Sinz / Panagiotis L Kastritis /   Abstract: The high abundance of most viruses in infected host cells benefits their structural characterization. However, endogenous viruses are present in low copy numbers and are therefore challenging to ...The high abundance of most viruses in infected host cells benefits their structural characterization. However, endogenous viruses are present in low copy numbers and are therefore challenging to investigate. Here, we retrieve cell extracts enriched with an endogenous virus, the yeast L-A virus. The determined cryo-EM structure discloses capsid-stabilizing cation-π stacking, widespread across viruses and within the Totiviridae, and an interplay of non-covalent interactions from ten distinct capsomere interfaces. The capsid-embedded mRNA decapping active site trench is supported by a constricting movement of two flexible opposite-facing loops. tRNA-loaded polysomes and other biomacromolecules, presumably mRNA, are found in virus proximity within the cell extract. Mature viruses participate in larger viral communities resembling their rare in-cell equivalents in terms of size, composition, and inter-virus distances. Our results collectively describe a 3D-architecture of a viral milieu, opening the door to cell-extract-based high-resolution structural virology. #1:  Journal: Biorxiv / Year: 2022 Journal: Biorxiv / Year: 2022Title: Delineating organizational principles of the endogenous L-A virus by cryo-EM and computational analysis of native cell extracts Authors: Schmidt L / Tuting C / Kyrilis FL / Hamdi F / Semchonok DA / Hause G / Meister A / Ihling C / Shah PNM / Stubbs MT / Sinz A / Stuart DI / Kastritis PL | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15214.map.gz emd_15214.map.gz | 59.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15214-v30.xml emd-15214-v30.xml emd-15214.xml emd-15214.xml | 21.1 KB 21.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15214_fsc.xml emd_15214_fsc.xml | 9.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_15214.png emd_15214.png | 77.5 KB | ||
| Masks |  emd_15214_msk_1.map emd_15214_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-15214.cif.gz emd-15214.cif.gz | 5 KB | ||
| Others |  emd_15214_additional_1.map.gz emd_15214_additional_1.map.gz emd_15214_half_map_1.map.gz emd_15214_half_map_1.map.gz emd_15214_half_map_2.map.gz emd_15214_half_map_2.map.gz | 40.3 MB 48.5 MB 48.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15214 http://ftp.pdbj.org/pub/emdb/structures/EMD-15214 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15214 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15214 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_15214.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15214.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Icosahedrally averaged capsid of the L-A helper virus from native Saccharomyces cerevisiae cell extract | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.177 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_15214_msk_1.map emd_15214_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
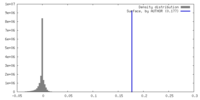
| Density Histograms |
-Additional map: Asymmetrically reconstructed inner density of the L-A helper...
| File | emd_15214_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Asymmetrically reconstructed inner density of the L-A helper virus revealing location of viral RNA | ||||||||||||
| Projections & Slices |
| ||||||||||||
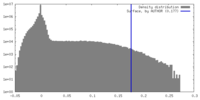
| Density Histograms |
-Half map: Icosahedrally averaged capsid of the L-A helper virus...
| File | emd_15214_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Icosahedrally averaged capsid of the L-A helper virus from native Saccharomyces cerevisiae cell extract - Half Map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
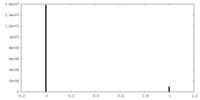
| Density Histograms |
-Half map: Icosahedrally averaged capsid of the L-A helper virus...
| File | emd_15214_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Icosahedrally averaged capsid of the L-A helper virus from native Saccharomyces cerevisiae cell extract - Half Map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
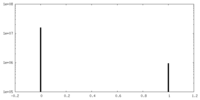
| Density Histograms |
- Sample components
Sample components
-Entire : Saccharomyces cerevisiae virus L-A
| Entire | Name:  Saccharomyces cerevisiae virus L-A Saccharomyces cerevisiae virus L-A |
|---|---|
| Components |
|
-Supramolecule #1: Saccharomyces cerevisiae virus L-A
| Supramolecule | Name: Saccharomyces cerevisiae virus L-A / type: virus / ID: 1 / Parent: 0 / Macromolecule list: #1 / NCBI-ID: 11008 / Sci species name: Saccharomyces cerevisiae virus L-A / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: SPECIES / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Component - Concentration: 200.0 mM / Component - Formula: CH3COONH4 / Component - Name: Ammoniumacetate Details: pH of the buffer was adjusted with NaOH buffer was filtered and sonicated |
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 15 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.04 kPa |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV / Details: blot force 2 and blot time 6 before plunging. |
| Details | heterogenous cell extract |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Temperature | Min: 77.0 K / Max: 118.0 K |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: INTEGRATING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 7 / Number real images: 10067 / Average exposure time: 3.61 sec. / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Calibrated magnification: 44067 / Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 45000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL |
|---|
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)