+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | In situ structure of HDCR filaments | ||||||||||||||||||
 Map data Map data | |||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
| Biological species |   Thermoanaerobacter kivui (bacteria) Thermoanaerobacter kivui (bacteria) | ||||||||||||||||||
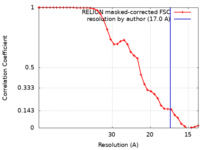
| Method | subtomogram averaging / cryo EM / Resolution: 17.0 Å | ||||||||||||||||||
 Authors Authors | Dietrich HM / Righetto RD / Kumar A / Wietrzynski W / Schuller SK / Trischler R / Wagner J / Schwarz FM / Engel BD / Mueller V / Schuller JM | ||||||||||||||||||
| Funding support |  Germany, European Union, 5 items Germany, European Union, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Nature / Year: 2022 Journal: Nature / Year: 2022Title: Membrane-anchored HDCR nanowires drive hydrogen-powered CO fixation. Authors: Helge M Dietrich / Ricardo D Righetto / Anuj Kumar / Wojciech Wietrzynski / Raphael Trischler / Sandra K Schuller / Jonathan Wagner / Fabian M Schwarz / Benjamin D Engel / Volker Müller / Jan M Schuller /   Abstract: Filamentous enzymes have been found in all domains of life, but the advantage of filamentation is often elusive. Some anaerobic, autotrophic bacteria have an unusual filamentous enzyme for CO ...Filamentous enzymes have been found in all domains of life, but the advantage of filamentation is often elusive. Some anaerobic, autotrophic bacteria have an unusual filamentous enzyme for CO fixation-hydrogen-dependent CO reductase (HDCR)-which directly converts H and CO into formic acid. HDCR reduces CO with a higher activity than any other known biological or chemical catalyst, and it has therefore gained considerable interest in two areas of global relevance: hydrogen storage and combating climate change by capturing atmospheric CO. However, the mechanistic basis of the high catalytic turnover rate of HDCR has remained unknown. Here we use cryo-electron microscopy to reveal the structure of a short HDCR filament from the acetogenic bacterium Thermoanaerobacter kivui. The minimum repeating unit is a hexamer that consists of a formate dehydrogenase (FdhF) and two hydrogenases (HydA2) bound around a central core of hydrogenase Fe-S subunits, one HycB3 and two HycB4. These small bacterial polyferredoxin-like proteins oligomerize through their C-terminal helices to form the backbone of the filament. By combining structure-directed mutagenesis with enzymatic analysis, we show that filamentation and rapid electron transfer through the filament enhance the activity of HDCR. To investigate the structure of HDCR in situ, we imaged T. kivui cells with cryo-electron tomography and found that HDCR filaments bundle into large ring-shaped superstructures attached to the plasma membrane. This supramolecular organization may further enhance the stability and connectivity of HDCR to form a specialized metabolic subcompartment within the cell. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15053.map.gz emd_15053.map.gz | 3.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15053-v30.xml emd-15053-v30.xml emd-15053.xml emd-15053.xml | 16.5 KB 16.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15053_fsc.xml emd_15053_fsc.xml | 3.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_15053.png emd_15053.png | 73.3 KB | ||
| Masks |  emd_15053_msk_1.map emd_15053_msk_1.map | 3.4 MB |  Mask map Mask map | |
| Others |  emd_15053_half_map_1.map.gz emd_15053_half_map_1.map.gz emd_15053_half_map_2.map.gz emd_15053_half_map_2.map.gz | 3.1 MB 3.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15053 http://ftp.pdbj.org/pub/emdb/structures/EMD-15053 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15053 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15053 | HTTPS FTP |
-Related structure data
| Related structure data |  7qv7C C: citing same article ( |
|---|---|
| EM raw data |  EMPIAR-11058 (Title: In situ cryo-electron tomography of T. kivui cells / Data size: 373.4 EMPIAR-11058 (Title: In situ cryo-electron tomography of T. kivui cells / Data size: 373.4 Data #1: Subtomograms of HDCR filament particles at bin2 [subtomograms] Data #2: Subtomograms of T. kivui ribosome particles at bin1 [subtomograms] Data #3: Denoised tomograms of T. kivui FIB-milled cells at bin4 [reconstructed volumes] Data #4: Tilt series of T. kivui FIB-milled cells - session 2020-10-22 [tilt series] Data #5: Tilt series of T. kivui FIB-milled cells - session 2021-02-12 [tilt series] Data #6: Unaligned raw frames of T. kivui FIB-milled cells tilt series - session 2020-10-22 [micrographs - multiframe] Data #7: Unaligned raw frames of T. kivui FIB-milled cells tilt series - session 2021-02-12 [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_15053.map.gz / Format: CCP4 / Size: 3.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15053.map.gz / Format: CCP4 / Size: 3.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 7.04 Å | ||||||||||||||||||||||||||||||||||||
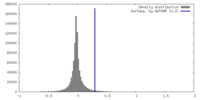
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_15053_msk_1.map emd_15053_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
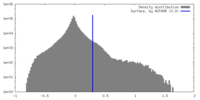
| Density Histograms |
-Half map: #1
| File | emd_15053_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
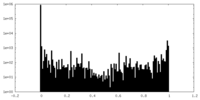
| Density Histograms |
-Half map: #2
| File | emd_15053_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
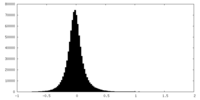
| Density Histograms |
- Sample components
Sample components
-Entire : Hydrogen-dependent CO2 reductase
| Entire | Name: Hydrogen-dependent CO2 reductase |
|---|---|
| Components |
|
-Supramolecule #1: Hydrogen-dependent CO2 reductase
| Supramolecule | Name: Hydrogen-dependent CO2 reductase / type: complex / Chimera: Yes / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:   Thermoanaerobacter kivui (bacteria) Thermoanaerobacter kivui (bacteria) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE-PROPANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Details | Dose-symmetric tilt-series were acquired (Hagen et al., 2017), starting at +10 degrees to match the pre-tilt of the lamella (i.e. from -50 to +70 degrees). |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 1.2 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 5.0 µm / Nominal defocus min: 3.5 µm / Nominal magnification: 42000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
|---|
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)