+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | cryo iDPC-STEM structure recorded with CSA 4.0 | |||||||||
 Map data Map data | cryo iDPC-STEM mao recorded with CSA 4.0 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | tobacco mosaic virus / RNA virus / VIRUS | |||||||||
| Function / homology | Tobacco mosaic virus-like, coat protein / Tobacco mosaic virus-like, coat protein superfamily / Virus coat protein (TMV like) / helical viral capsid / structural molecule activity / identical protein binding / Capsid protein Function and homology information Function and homology information | |||||||||
| Biological species |  Tobacco mosaic virus (vulgare) / Tobacco mosaic virus (vulgare) /   Tobacco mosaic virus Tobacco mosaic virus | |||||||||
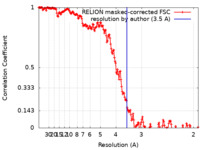
| Method | helical reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Sachse C / Leidl ML | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Methods / Year: 2022 Journal: Nat Methods / Year: 2022Title: Single-particle cryo-EM structures from iDPC-STEM at near-atomic resolution. Authors: Ivan Lazić / Maarten Wirix / Max Leo Leidl / Felix de Haas / Daniel Mann / Maximilian Beckers / Evgeniya V Pechnikova / Knut Müller-Caspary / Ricardo Egoavil / Eric G T Bosch / Carsten Sachse /   Abstract: In electron cryomicroscopy (cryo-EM), molecular images of vitrified biological samples are obtained by conventional transmission microscopy (CTEM) using large underfocuses and subsequently ...In electron cryomicroscopy (cryo-EM), molecular images of vitrified biological samples are obtained by conventional transmission microscopy (CTEM) using large underfocuses and subsequently computationally combined into a high-resolution three-dimensional structure. Here, we apply scanning transmission electron microscopy (STEM) using the integrated differential phase contrast mode also known as iDPC-STEM to two cryo-EM test specimens, keyhole limpet hemocyanin (KLH) and tobacco mosaic virus (TMV). The micrographs show complete contrast transfer to high resolution and enable the cryo-EM structure determination for KLH at 6.5 Å resolution, as well as for TMV at 3.5 Å resolution using single-particle reconstruction methods, which share identical features with maps obtained by CTEM of a previously acquired same-sized TMV data set. These data show that STEM imaging in general, and in particular the iDPC-STEM approach, can be applied to vitrified single-particle specimens to determine near-atomic resolution cryo-EM structures of biological macromolecules. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13781.map.gz emd_13781.map.gz | 22.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13781-v30.xml emd-13781-v30.xml emd-13781.xml emd-13781.xml | 15.8 KB 15.8 KB | Display Display |  EMDB header EMDB header |
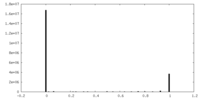
| FSC (resolution estimation) |  emd_13781_fsc.xml emd_13781_fsc.xml | 17.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_13781.png emd_13781.png | 136.7 KB | ||
| Masks |  emd_13781_msk_1.map emd_13781_msk_1.map | 476.8 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-13781.cif.gz emd-13781.cif.gz | 5.3 KB | ||
| Others |  emd_13781_half_map_1.map.gz emd_13781_half_map_1.map.gz emd_13781_half_map_2.map.gz emd_13781_half_map_2.map.gz | 381.7 MB 381.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13781 http://ftp.pdbj.org/pub/emdb/structures/EMD-13781 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13781 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13781 | HTTPS FTP |
-Related structure data
| Related structure data |  7q2rMC  7q22C  7q23C  7q2qC  7q2sC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_13781.map.gz / Format: CCP4 / Size: 143.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13781.map.gz / Format: CCP4 / Size: 143.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cryo iDPC-STEM mao recorded with CSA 4.0 | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.98 Å | ||||||||||||||||||||||||||||||||||||
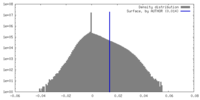
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_13781_msk_1.map emd_13781_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half-map1
| File | emd_13781_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half-map1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half-map2
| File | emd_13781_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half-map2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Tobacco mosaic virus
| Entire | Name:   Tobacco mosaic virus Tobacco mosaic virus |
|---|---|
| Components |
|
-Supramolecule #1: Tobacco mosaic virus
| Supramolecule | Name: Tobacco mosaic virus / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 12242 / Sci species name: Tobacco mosaic virus / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:   Tobacco mosaic virus Tobacco mosaic virus |
| Molecular weight | Theoretical: 131 kDa/nm |
-Macromolecule #1: Capsid protein
| Macromolecule | Name: Capsid protein / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Tobacco mosaic virus (vulgare) / Strain: vulgare Tobacco mosaic virus (vulgare) / Strain: vulgare |
| Molecular weight | Theoretical: 17.091998 KDa |
| Sequence | String: SYSITTPSQF VFLSSAWADP IELINLCTNA LGNQFQTQQA RTVVQRQFSE VWKPSPQVTV RFPDSDFKVY RYNAVLDPLV TALLGAFDT RNRIIEVENQ ANPTTAETLD ATRRVDDATV AIRSAINNLI VELIRGTGSY NRSSFESSSG LVWT UniProtKB: Capsid protein |
-Macromolecule #2: RNA (5'-R(P*GP*AP*A)-3')
| Macromolecule | Name: RNA (5'-R(P*GP*AP*A)-3') / type: rna / ID: 2 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  Tobacco mosaic virus (vulgare) Tobacco mosaic virus (vulgare) |
| Molecular weight | Theoretical: 958.66 Da |
| Sequence | String: GAA |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | helical array |
- Sample preparation
Sample preparation
| Concentration | 90 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV Details: blot force of +10 and a duration for blotting of 10 seconds. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Details | iDPC-STEM |
| Image recording | Film or detector model: OTHER / Average electron dose: 35.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: OTHER / Nominal defocus max: 0.0 µm / Nominal defocus min: 0.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
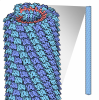










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)