+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10160 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | In Situ Core-Signalling Unit of E. coli Chemoreceptor Array | |||||||||
 Map data Map data | None | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 16.0 Å | |||||||||
 Authors Authors | Burt A / Desfosses A / Gutsche I | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2020 Journal: Nat Commun / Year: 2020Title: Complete structure of the chemosensory array core signalling unit in an E. coli minicell strain. Authors: Alister Burt / C Keith Cassidy / Peter Ames / Maria Bacia-Verloop / Megghane Baulard / Karine Huard / Zaida Luthey-Schulten / Ambroise Desfosses / Phillip J Stansfeld / William Margolin / ...Authors: Alister Burt / C Keith Cassidy / Peter Ames / Maria Bacia-Verloop / Megghane Baulard / Karine Huard / Zaida Luthey-Schulten / Ambroise Desfosses / Phillip J Stansfeld / William Margolin / John S Parkinson / Irina Gutsche /    Abstract: Motile bacteria sense chemical gradients with transmembrane receptors organised in supramolecular signalling arrays. Understanding stimulus detection and transmission at the molecular level requires ...Motile bacteria sense chemical gradients with transmembrane receptors organised in supramolecular signalling arrays. Understanding stimulus detection and transmission at the molecular level requires precise structural characterisation of the array building block known as a core signalling unit. Here we introduce an Escherichia coli strain that forms small minicells possessing extended and highly ordered chemosensory arrays. We use cryo-electron tomography and subtomogram averaging to provide a three-dimensional map of a complete core signalling unit, with visible densities corresponding to the HAMP and periplasmic domains. This map, combined with previously determined high resolution structures and molecular dynamics simulations, yields a molecular model of the transmembrane core signalling unit and enables spatial localisation of its individual domains. Our work thus offers a solid structural basis for the interpretation of a wide range of existing data and the design of further experiments to elucidate signalling mechanisms within the core signalling unit and larger array. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10160.map.gz emd_10160.map.gz | 4.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10160-v30.xml emd-10160-v30.xml emd-10160.xml emd-10160.xml | 16.9 KB 16.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_10160_fsc.xml emd_10160_fsc.xml | 4.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_10160.png emd_10160.png | 55.5 KB | ||
| Masks |  emd_10160_msk_1.map emd_10160_msk_1.map emd_10160_msk_2.map emd_10160_msk_2.map | 8 MB 8 MB |  Mask map Mask map | |
| Others |  emd_10160_additional.map.gz emd_10160_additional.map.gz emd_10160_additional_1.map.gz emd_10160_additional_1.map.gz emd_10160_half_map_1.map.gz emd_10160_half_map_1.map.gz emd_10160_half_map_2.map.gz emd_10160_half_map_2.map.gz | 4.5 MB 4.5 MB 7.4 MB 7.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10160 http://ftp.pdbj.org/pub/emdb/structures/EMD-10160 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10160 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10160 | HTTPS FTP |
-Related structure data
| Similar structure data | |
|---|---|
| EM raw data |  EMPIAR-10364 (Title: Cryo-electron tomography of E. coli minicells / Data size: 141.7 EMPIAR-10364 (Title: Cryo-electron tomography of E. coli minicells / Data size: 141.7 Data #1: Unaligned multi-frame micrographs from tilt series containing WM4196 minicells [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_10160.map.gz / Format: CCP4 / Size: 8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10160.map.gz / Format: CCP4 / Size: 8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.48 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_10160_msk_1.map emd_10160_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
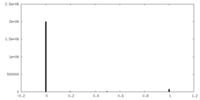
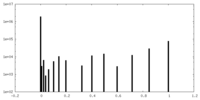
| Density Histograms |
-Mask #2
| File |  emd_10160_msk_2.map emd_10160_msk_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Local resolution filtered map centered on 3fold axis...
| File | emd_10160_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Local resolution filtered map centered on 3fold axis of chemoreceptor array. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Local resolution filtered map centered on 3fold axis...
| File | emd_10160_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Local resolution filtered map centered on 3fold axis of chemoreceptor array. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: even half map.
| File | emd_10160_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | even half map. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: odd half map.
| File | emd_10160_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | odd half map. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Optimised E. coli minicell strain WM4196 containing native-state ...
| Entire | Name: Optimised E. coli minicell strain WM4196 containing native-state chemoreceptor arrays. |
|---|---|
| Components |
|
-Supramolecule #1: Optimised E. coli minicell strain WM4196 containing native-state ...
| Supramolecule | Name: Optimised E. coli minicell strain WM4196 containing native-state chemoreceptor arrays. type: cell / ID: 1 / Parent: 0 Details: Minicells isolated from WM4196 cell culture by differential centrifugation. |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Grid | Model: Quantifoil R3.5/1 / Material: COPPER/RHODIUM / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR / Details: 25 mA |
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Phase plate: VOLTA PHASE PLATE |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Frames/image: 1-5 / Number grids imaged: 1 / Average electron dose: 1.0 e/Å2 Details: Images were collected in movie mode, 5 frames per movie. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus min: 0.0002 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)