+ Open data
Open data
- Basic information
Basic information
| Entry | 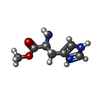 Database: PDB chemical components / ID: PVH Database: PDB chemical components / ID: PVH | ||
|---|---|---|---|
| Name | Name: | Comment | inhibitor*YM | |
-Chemical information
| Composition |  | ||||||
|---|---|---|---|---|---|---|---|
| Others | Type: L-PEPTIDE LINKING / PDB classification: ATOMP / One letter code: H / Three letter code: PVH / Parent comp.: HIS | ||||||
| History |
| ||||||
 External links External links |  UniChem / UniChem /  ChemSpider / ChemSpider /  PubChem / PubChem /  Wikipedia search / Wikipedia search /  Google search Google search |
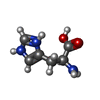
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
-Details
-SMILES
| ACDLabs 10.04 | | CACTVS 3.341 | OpenEye OEToolkits 1.5.0 | |
|---|
-SMILES CANONICAL
| CACTVS 3.341 | | OpenEye OEToolkits 1.5.0 | |
|---|
-InChI
| InChI 1.03 |
|---|
-InChIKey
| InChI 1.03 |
|---|
-SYSTEMATIC NAME
| ACDLabs 10.04 | | OpenEye OEToolkits 1.5.0 | |
|---|
-PDB entries
Showing all 3 items

PDB-1ibv: 
STRUCTURE OF THE D53,54N MUTANT OF HISTIDINE DECARBOXYLASE BOUND WITH HISTIDINE METHYL ESTER AT-170 C

PDB-1ibw: 
STRUCTURE OF THE D53,54N MUTANT OF HISTIDINE DECARBOXYLASE BOUND WITH HISTIDINE METHYL ESTER AT 25 C

PDB-4e1o: 
Human histidine decarboxylase complex with Histidine methyl ester (HME)
 Movie
Movie Controller
Controller