[English] 日本語
 Yorodumi
Yorodumi- ChemComp-0HZ: amino({3-[(3S,8aS)-1,4-dioxooctahydropyrrolo[1,2-a]pyrazin-3-yl]p... -
+ Open data
Open data
- Basic information
Basic information
| Entry | 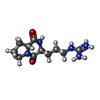 Database: PDB chemical components / ID: 0HZ Database: PDB chemical components / ID: 0HZ |
|---|---|
| Name | Name: Synonyms: CI-4; [cyclo-(l-Arg-d-Pro)] |
-Chemical information
| Composition |  | ||||||
|---|---|---|---|---|---|---|---|
| Others | Type: peptide-like / PDB classification: HETAIN / Three letter code: 0HZ / Ideal coordinates details: Corina / Model coordinates PDB-ID: 1O6I / Model coordinates details: not provided / Subcomponent: DPR, ARG | ||||||
| History |
| ||||||
 External links External links |  UniChem / UniChem /  ChemSpider / ChemSpider /  PubChem / PubChem /  Wikipedia search / Wikipedia search /  Google search Google search |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
-Details
-SMILES
| ACDLabs 10.04 | | CACTVS 3.341 | |
|---|
-SMILES CANONICAL
| CACTVS 3.341 |
|---|
-InChI
| InChI 1.03 |
|---|
-InChIKey
| InChI 1.03 |
|---|
-SYSTEMATIC NAME
| ACDLabs 10.04 | | OpenEye OEToolkits 1.5.0 | [[ | |
|---|
-PDB entries
Showing all 5 items

PDB-1o6i: 
Chitinase B from Serratia marcescens complexed with the catalytic intermediate mimic cyclic dipeptide CI4.

PDB-5yvk: 
Crystal structure of a cyclase Famc1 from Fischerella ambigua UTEX 1903

PDB-5z53: 
Crystal structure of a cyclase Filc from Fischerella sp. in complex with cyclo-L-Arg-D-Pro

PDB-5z54: 
Crystal structure of a cyclase Hpiu5 from Fischerella sp. ATCC 43239 in complex with cyclo-L-Arg-D-Pro

PDB-6j03: 
Crystal structure of a cyclase mutant in apo form
 Movie
Movie Controller
Controller