[English] 日本語
 Yorodumi
Yorodumi- ChemComp-AVN: (2S)-AMINO[(5S)-3-CHLORO-4,5-DIHYDROISOXAZOL-5-YL]ACETIC ACID -
+ Open data
Open data
- Basic information
Basic information
| Entry | 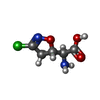 Database: PDB chemical components / ID: AVN Database: PDB chemical components / ID: AVN |
|---|---|
| Name | Name: ( Synonyms: ACIVICIN |
-Chemical information
| Composition |  | ||||||
|---|---|---|---|---|---|---|---|
| Others | Type: L-PEPTIDE LINKING / PDB classification: ATOMP / One letter code: X / Three letter code: AVN / Ideal coordinates details: Corina / Model coordinates PDB-ID: 2Z8K | ||||||
| History |
| ||||||
 External links External links |  UniChem / UniChem /  ChemSpider / ChemSpider /  Brenda Brenda     / /  ChEBI / ChEBI /  ChEMBL / ChEMBL /  CompTox / CompTox /  DrugBank / DrugBank /  PubChem PubChem  / /  SureChEMBL / SureChEMBL /  Wikipedia search / Wikipedia search /  Google search Google search |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
-Details
-SMILES
| ACDLabs 10.04 | | CACTVS 3.341 | OpenEye OEToolkits 1.5.0 | |
|---|
-SMILES CANONICAL
| CACTVS 3.341 | | OpenEye OEToolkits 1.5.0 | |
|---|
-InChI
| InChI 1.03 |
|---|
-InChIKey
| InChI 1.03 |
|---|
-SYSTEMATIC NAME
| ACDLabs 10.04 | (| OpenEye OEToolkits 1.5.0 | ( | |
|---|
-PDB entries
Showing all 4 items

PDB-2z8k: 
Crystal Structure of Escherichia coli gamma-Glutamyltranspeptidase in Complex with Acivicin

PDB-3fnm: 
Crystal structure of acivicin-inhibited gamma-glutamyltranspeptidase reveals critical roles for its C-terminus in autoprocessing and catalysis

PDB-3whs: 
Crystal structure of Bacillus subtilis gamma-glutamyltranspeptidase in complex with acivicin

PDB-5xlu: 
High Resolution Crystal Structure of Bacillus Licheniformis Gamma Glutamyl Transpeptidase with Acivicin
 Movie
Movie Controller
Controller


