[English] 日本語
 Yorodumi
Yorodumi- EMDB-7822: Cryo-EM structure of the zebrafish TRPM2 channel in the presence ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-7822 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the zebrafish TRPM2 channel in the presence of Ca2+ | |||||||||
 Map data Map data | Single-particle cryo-EM reconstruction of TRPM2 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | warmth sensor / redox sensor / calcium-permeable ion channel / ion channel / TRANSPORT PROTEIN | |||||||||
| Biological species |  | |||||||||
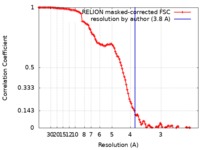
| Method | single particle reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||
 Authors Authors | Yin Y / Wu M | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2019 Journal: Nat Commun / Year: 2019Title: Visualizing structural transitions of ligand-dependent gating of the TRPM2 channel. Authors: Ying Yin / Mengyu Wu / Allen L Hsu / William F Borschel / Mario J Borgnia / Gabriel C Lander / Seok-Yong Lee /  Abstract: The transient receptor potential melastatin 2 (TRPM2) channel plays a key role in redox sensation in many cell types. Channel activation requires binding of both ADP-ribose (ADPR) and Ca. The ...The transient receptor potential melastatin 2 (TRPM2) channel plays a key role in redox sensation in many cell types. Channel activation requires binding of both ADP-ribose (ADPR) and Ca. The recently published TRPM2 structures from Danio rerio in the ligand-free and the ADPR/Ca-bound conditions represent the channel in closed and open states, which uncovered substantial tertiary and quaternary conformational rearrangements. However, it is unclear how these rearrangements are achieved within the tetrameric channel during channel gating. Here we report the cryo-electron microscopy structures of Danio rerio TRPM2 in the absence of ligands, in complex with Ca alone, and with both ADPR and Ca, resolved to ~4.3 Å, ~3.8 Å, and ~4.2 Å, respectively. In contrast to the published results, our studies capture ligand-bound TRPM2 structures in two-fold symmetric intermediate states, offering a glimpse of the structural transitions that bridge the closed and open conformations. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_7822.map.gz emd_7822.map.gz | 113 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-7822-v30.xml emd-7822-v30.xml emd-7822.xml emd-7822.xml | 18.5 KB 18.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_7822_fsc.xml emd_7822_fsc.xml | 11.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_7822.png emd_7822.png | 246.1 KB | ||
| Filedesc metadata |  emd-7822.cif.gz emd-7822.cif.gz | 7 KB | ||
| Others |  emd_7822_half_map_1.map.gz emd_7822_half_map_1.map.gz emd_7822_half_map_2.map.gz emd_7822_half_map_2.map.gz | 96.4 MB 96.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7822 http://ftp.pdbj.org/pub/emdb/structures/EMD-7822 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7822 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7822 | HTTPS FTP |
-Related structure data
| Related structure data |  6d73MC  6pkvC  6pkwC  6pkxC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_7822.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_7822.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Single-particle cryo-EM reconstruction of TRPM2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.15 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: Half 1
| File | emd_7822_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half 1 | ||||||||||||
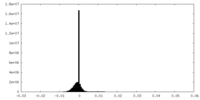
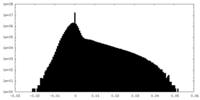
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half 2
| File | emd_7822_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half 2 | ||||||||||||
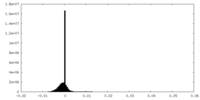
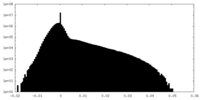
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Transient receptor potential cation channel subfamily M member 2 ...
| Entire | Name: Transient receptor potential cation channel subfamily M member 2 (TRPM2) |
|---|---|
| Components |
|
-Supramolecule #1: Transient receptor potential cation channel subfamily M member 2 ...
| Supramolecule | Name: Transient receptor potential cation channel subfamily M member 2 (TRPM2) type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Transient receptor potential cation channel, subfamily M
| Macromolecule | Name: Transient receptor potential cation channel, subfamily M type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 167.608625 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MALGTSGVKI HPNGNSNQLG VQLENVKLTS LFKKLDKRCS LASWIKENIK KKECCFYVED GREGICKCGY PKVQHCDEAI KPEDYMGEQ WDKHRHVRET PTDAFGDISF GGLGQKTGKY VRVSSDTSCE NLYQLMTEQW KLRSPNLLIS VTGGAKNFYI K THLKDKFR ...String: MALGTSGVKI HPNGNSNQLG VQLENVKLTS LFKKLDKRCS LASWIKENIK KKECCFYVED GREGICKCGY PKVQHCDEAI KPEDYMGEQ WDKHRHVRET PTDAFGDISF GGLGQKTGKY VRVSSDTSCE NLYQLMTEQW KLRSPNLLIS VTGGAKNFYI K THLKDKFR RGLIKVAQTT GAWILTGGTH AGVMKHVGMA VRDYTLSSGS MEGQIVVIGV APWGVIHNRS TLIHPEGRFP AY YSLDEQG QGRLSCLDIN HTHFLLVDDG TQGHYGVEIE LRARLEKLIS KLSLGNRESG VTIPVVCVVL DGGPGTLNTI YNS MLNHTP CVVLEGSGRL ADVIAHVASV PVSKVTMALI NRLLKRFFMQ EYKNFTELQI IEWTKKIQDI LRMPHLLTVF RIDE DKNYD VDVAILQALL KASRSDEHAG RHCWERQLEL AVAWNRVDIA ESEIFTEESQ WTSSDLHPAM FSALVGDKPE FVRLL LENG VCVREFLERE ETLCELYSHL PSCFFLRKLA KRVQGGKMRR GQEPLPGSRK VCLSHVSEEV RHLLGSFTQP LYIASR YKP TKDDVRLKVP SKGALDLPCS GEEWSADTVW DPGRDLFLWA VVQNNRELAE IGWEQCRDCI AAALAASKIL RKLAQES GE DDSEEATEML ELANHYEKQA IGVFSECHSW DAQRAQKLLI RISPSWGRST CLWLALEAHD KSFIAHSGVQ ALLTQIWC G ELSVDNPHWK VLLCMIFFPL IYTGFLTFRR DEDIQRQAER TEQQKLAMES VFAGQSDGKI KRHLRGFSQK SELKPLNCS SRLMSFLKSP QVKFYWNIAS YFGFLWLFAV VLMIDFQTSP SWRELLLYVW LTSLVCEEIR QLYHDFDGSG FRRKAKMYIK DLWNILDVL SIVLFIAGLI CRLQASDTVF YIGKVILCID FIIFCLRLMA IFSISRTLGP KIIIVRRMML DLFFFMFLLS I WVVAYGVA KQGILIENEE RLNWIIRGAV YEPYITIFGN FPTNIDNTLF DISSCSVNAS DPLKPKCPML NADNTPVFPE WL TIMMLCV YLLFANILLL NLLIAIFNYT FQEVQDNTDT IWKFQRYELI KEYHSRPALP PPFILLSHLI LFIRGVFLRD LPQ RHKNFR QELEQTEEEE LLSWEAYMKD NYLASTRQDE SQSVEHRIHD TAEKVGAMSE LLEREQEMV(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)DEEAPHM FARQLQYPDS TVRRFPVPEE KVSWEVNFSP YQPPVYNQQD SSESDTSALD KHRNPGGR T GIRGKGALNT LGPNHILHPI FTRWRDAEHK VLEFLAVWED AEKRWALLGG PAQPDEPLAQ VLERILGKKL NEKTKTLLK AGEEVYKGYV DDSRNTDNAW VETSIITLHC DKNTPLMADL NHMVESSLSS HQPLQWREVS SDACRCSYQR EALRQIAHHH NTYFSNSLE VLFQGPDYKD DDDKAHHHHH HHHHH |
-Macromolecule #2: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 2 / Number of copies: 4 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Grid | Model: Quantifoil, UltrAuFoil, R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Pretreatment - Type: PLASMA CLEANING |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: HOMEMADE PLUNGER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 3710 pixel / Digitization - Dimensions - Height: 3838 pixel / Digitization - Frames/image: 1-64 / Number real images: 3039 / Average exposure time: 16.0 sec. / Average electron dose: 63.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.2 µm / Nominal magnification: 36000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Overall B value: 100 |
|---|---|
| Output model |  PDB-6d73: |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)