[English] 日本語
 Yorodumi
Yorodumi- EMDB-45008: Paired Helical Filaments purified from Down Syndrome individual b... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Paired Helical Filaments purified from Down Syndrome individual brain tissue applied to graphene oxide antibody affinity grids | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Amyloid / filament / neurodegeneration / Down syndrome / PROTEIN FIBRIL | |||||||||
| Function / homology |  Function and homology information Function and homology informationplus-end-directed organelle transport along microtubule / histone-dependent DNA binding / neurofibrillary tangle assembly / positive regulation of diacylglycerol kinase activity / axonal transport / negative regulation of establishment of protein localization to mitochondrion / neurofibrillary tangle / positive regulation of protein localization to synapse / microtubule lateral binding / tubulin complex ...plus-end-directed organelle transport along microtubule / histone-dependent DNA binding / neurofibrillary tangle assembly / positive regulation of diacylglycerol kinase activity / axonal transport / negative regulation of establishment of protein localization to mitochondrion / neurofibrillary tangle / positive regulation of protein localization to synapse / microtubule lateral binding / tubulin complex / phosphatidylinositol bisphosphate binding / main axon / negative regulation of kinase activity / regulation of long-term synaptic depression / negative regulation of tubulin deacetylation / generation of neurons / rRNA metabolic process / internal protein amino acid acetylation / regulation of chromosome organization / regulation of mitochondrial fission / axonal transport of mitochondrion / intracellular distribution of mitochondria / axon development / central nervous system neuron development / regulation of microtubule polymerization / apolipoprotein binding / microtubule polymerization / lipoprotein particle binding / minor groove of adenine-thymine-rich DNA binding / dynactin binding / glial cell projection / negative regulation of mitochondrial membrane potential / protein polymerization / axolemma / negative regulation of mitochondrial fission / Caspase-mediated cleavage of cytoskeletal proteins / regulation of microtubule polymerization or depolymerization / positive regulation of axon extension / regulation of microtubule cytoskeleton organization / Activation of AMPK downstream of NMDARs / regulation of cellular response to heat / positive regulation of protein localization / cytoplasmic microtubule organization / stress granule assembly / supramolecular fiber organization / regulation of calcium-mediated signaling / axon cytoplasm / somatodendritic compartment / positive regulation of microtubule polymerization / synapse assembly / cellular response to brain-derived neurotrophic factor stimulus / phosphatidylinositol binding / nuclear periphery / cellular response to nerve growth factor stimulus / positive regulation of superoxide anion generation / protein phosphatase 2A binding / regulation of autophagy / astrocyte activation / response to lead ion / microglial cell activation / synapse organization / Hsp90 protein binding / protein homooligomerization / PKR-mediated signaling / regulation of synaptic plasticity / : / memory / SH3 domain binding / microtubule cytoskeleton organization / cytoplasmic ribonucleoprotein granule / cellular response to reactive oxygen species / microtubule cytoskeleton / neuron projection development / cell-cell signaling / single-stranded DNA binding / protein-folding chaperone binding / actin binding / protein-macromolecule adaptor activity / cellular response to heat / double-stranded DNA binding / growth cone / cell body / microtubule binding / microtubule / sequence-specific DNA binding / amyloid fibril formation / dendritic spine / learning or memory / nuclear speck / neuron projection / membrane raft / axon / negative regulation of gene expression / neuronal cell body / DNA damage response / dendrite / protein kinase binding / enzyme binding / mitochondrion / DNA binding Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Tse E / Ghosh U / Condello C / Southworth D | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Acta Neuropathol Commun / Year: 2024 Journal: Acta Neuropathol Commun / Year: 2024Title: Cryo-EM structures reveal tau filaments from Down syndrome adopt Alzheimer's disease fold. Authors: Ujjayini Ghosh / Eric Tse / Hyunjun Yang / Marie Shi / Christoffer D Caro / Feng Wang / Gregory E Merz / Stanley B Prusiner / Daniel R Southworth / Carlo Condello /  Abstract: Down syndrome (DS) is a common genetic condition caused by trisomy of chromosome 21. Among their complex clinical features, including musculoskeletal, neurological, and cardiovascular disabilities, ...Down syndrome (DS) is a common genetic condition caused by trisomy of chromosome 21. Among their complex clinical features, including musculoskeletal, neurological, and cardiovascular disabilities, individuals with DS have an increased risk of developing progressive dementia and early-onset Alzheimer's disease (AD). This dementia is attributed to the increased gene dosage of the amyloid-β (Aβ) precursor protein gene, the formation of self-propagating Aβ and tau prion conformers, and the deposition of neurotoxic Aβ plaques and tau neurofibrillary tangles. Tau amyloid fibrils have previously been established to adopt many distinct conformations across different neurodegenerative conditions. Here, we report the characterization of brain samples from four DS cases spanning 36-63 years of age by spectral confocal imaging with conformation-specific dyes and cryo-electron microscopy (cryo-EM) to determine structures of isolated tau fibrils. High-resolution structures revealed paired helical filament (PHF) and straight filament (SF) conformations of tau that were identical to those determined from AD cases. The PHFs and SFs are made of two C-shaped protofilaments, each containing a cross-β/β-helix motif. Similar to filaments from AD cases, most filaments from the DS cases adopted the PHF form, while a minority (approximately 20%) formed SFs. Samples from the youngest individual with no documented dementia had sparse tau deposits. To isolate tau for cryo-EM from this challenging sample we used a novel affinity-grid method involving a graphene oxide surface derivatized with anti-tau antibodies. This method improved isolation and revealed that primarily tau PHFs and a minor population of chronic traumatic encephalopathy type II-like filaments were present in this youngest case. These findings expand the similarities between AD and DS to the molecular level, providing insight into their related pathologies and the potential for targeting common tau filament folds by small-molecule therapeutics and diagnostics. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_45008.map.gz emd_45008.map.gz | 8.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-45008-v30.xml emd-45008-v30.xml emd-45008.xml emd-45008.xml | 14.5 KB 14.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_45008.png emd_45008.png | 75.9 KB | ||
| Filedesc metadata |  emd-45008.cif.gz emd-45008.cif.gz | 5.1 KB | ||
| Others |  emd_45008_half_map_1.map.gz emd_45008_half_map_1.map.gz emd_45008_half_map_2.map.gz emd_45008_half_map_2.map.gz | 65.3 MB 65.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-45008 http://ftp.pdbj.org/pub/emdb/structures/EMD-45008 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-45008 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-45008 | HTTPS FTP |
-Validation report
| Summary document |  emd_45008_validation.pdf.gz emd_45008_validation.pdf.gz | 798.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_45008_full_validation.pdf.gz emd_45008_full_validation.pdf.gz | 798.5 KB | Display | |
| Data in XML |  emd_45008_validation.xml.gz emd_45008_validation.xml.gz | 12.8 KB | Display | |
| Data in CIF |  emd_45008_validation.cif.gz emd_45008_validation.cif.gz | 14.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-45008 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-45008 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-45008 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-45008 | HTTPS FTP |
-Related structure data
| Related structure data |  9bxqMC  9bxiC  9bxoC  9bxrC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_45008.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_45008.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.834 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_45008_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
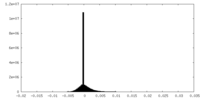
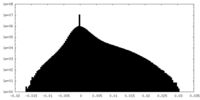
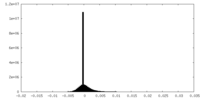
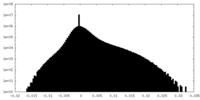
| Density Histograms |
-Half map: #1
| File | emd_45008_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Paired Helical Filaments purified from Down Syndrome individual b...
| Entire | Name: Paired Helical Filaments purified from Down Syndrome individual brain tissue applied to graphene oxide antibody affinity grids |
|---|---|
| Components |
|
-Supramolecule #1: Paired Helical Filaments purified from Down Syndrome individual b...
| Supramolecule | Name: Paired Helical Filaments purified from Down Syndrome individual brain tissue applied to graphene oxide antibody affinity grids type: tissue / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Microtubule-associated protein tau
| Macromolecule | Name: Microtubule-associated protein tau / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 8.370578 KDa |
| Sequence | String: GSVQIVYKPV DLSKVTSKCG SLGNIHHKPG GGQVEVKSEK LDFKDRVQSK IGSLDNITHV PGGGNKKIET HKLTFRE UniProtKB: Microtubule-associated protein tau |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
| Details | Sarkosyl extraction of insoluble tau from fresh-frozen frontal cortex samples from Down Syndrome individuals |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 56.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 2.37 Å Applied symmetry - Helical parameters - Δ&Phi: 179.5 ° Applied symmetry - Helical parameters - Axial symmetry: C1 (asymmetric) Resolution.type: BY AUTHOR / Resolution: 3.1 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 79599 |
|---|---|
| Startup model | Type of model: INSILICO MODEL / In silico model: relion_helix_inimodel2d Details: selected class averages with similar views along filament length |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)