+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of HMPV (MPV-2cREKR) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Prefusion / HMPV / Cryo-EM / VIRUS / VIRAL PROTEIN | |||||||||
| Function / homology | Precursor fusion glycoprotein F0, Paramyxoviridae / Fusion glycoprotein F0 / symbiont entry into host cell / fusion of virus membrane with host plasma membrane / viral envelope / host cell plasma membrane / virion membrane / membrane / Fusion glycoprotein F0 Function and homology information Function and homology information | |||||||||
| Biological species |  Human metapneumovirus Human metapneumovirus | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.33 Å | |||||||||
 Authors Authors | Yu X / Langedijk JPM | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Efficacious human metapneumovirus vaccine based on AI-guided engineering of a closed prefusion trimer Authors: Bakkers MJG / Ritschel T / Tiemessen M / Dijkman J / Zuffiano AA / Yu X / Overveld D / Le L / Voorzaat R / Haaren M / Man M / Sharma S / Tamara S / Fits L / Zahn R / Juraszek J / Langedijk JPM | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_43517.map.gz emd_43517.map.gz | 118 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-43517-v30.xml emd-43517-v30.xml emd-43517.xml emd-43517.xml | 14.6 KB 14.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_43517.png emd_43517.png | 39.4 KB | ||
| Filedesc metadata |  emd-43517.cif.gz emd-43517.cif.gz | 5.6 KB | ||
| Others |  emd_43517_half_map_1.map.gz emd_43517_half_map_1.map.gz emd_43517_half_map_2.map.gz emd_43517_half_map_2.map.gz | 116.1 MB 116.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-43517 http://ftp.pdbj.org/pub/emdb/structures/EMD-43517 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43517 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43517 | HTTPS FTP |
-Validation report
| Summary document |  emd_43517_validation.pdf.gz emd_43517_validation.pdf.gz | 1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_43517_full_validation.pdf.gz emd_43517_full_validation.pdf.gz | 1 MB | Display | |
| Data in XML |  emd_43517_validation.xml.gz emd_43517_validation.xml.gz | 14 KB | Display | |
| Data in CIF |  emd_43517_validation.cif.gz emd_43517_validation.cif.gz | 16.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43517 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43517 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43517 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43517 | HTTPS FTP |
-Related structure data
| Related structure data |  8vt3MC  8vt2C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_43517.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_43517.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.91 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_43517_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
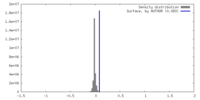
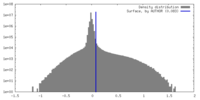
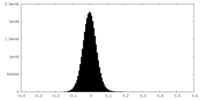
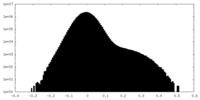
| Density Histograms |
-Half map: #1
| File | emd_43517_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
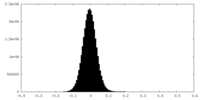
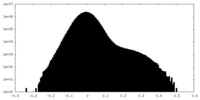
| Density Histograms |
- Sample components
Sample components
-Entire : Prefusion of HMPV (MPV-2cREKR) trimer complex
| Entire | Name: Prefusion of HMPV (MPV-2cREKR) trimer complex |
|---|---|
| Components |
|
-Supramolecule #1: Prefusion of HMPV (MPV-2cREKR) trimer complex
| Supramolecule | Name: Prefusion of HMPV (MPV-2cREKR) trimer complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Human metapneumovirus Human metapneumovirus |
-Macromolecule #1: Prefusion of HMPV (MPV-2cREKR), N-fragment
| Macromolecule | Name: Prefusion of HMPV (MPV-2cREKR), N-fragment / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Human metapneumovirus Human metapneumovirus |
| Molecular weight | Theoretical: 8.296239 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GLKESYLEES CSTITEGYLS VLRTGWYTNV FTLEVGDVEN LTCSDGPSLI KTELDLTKSA LRELKTVSAD QLARE UniProtKB: Fusion glycoprotein F0 |
-Macromolecule #2: Prefusion of HMPV (MPV-2cREKR), C-fragment
| Macromolecule | Name: Prefusion of HMPV (MPV-2cREKR), C-fragment / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Human metapneumovirus Human metapneumovirus |
| Molecular weight | Theoretical: 41.865828 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: FVLGAIALGR ATAAAVTAGV AIAKTIRLES EVTAIKNALK TTNEAVSTLG NGVRVLATAV RELKDFVSKN LTRAINKNKC DIDDLKMAV SFSQFNRRFL NVVRQFSENA GITPAISLDL MTDAELARAI SNMPTSAGQI KLMLENRAMV RRKGFGILIG V YGSSVIYM ...String: FVLGAIALGR ATAAAVTAGV AIAKTIRLES EVTAIKNALK TTNEAVSTLG NGVRVLATAV RELKDFVSKN LTRAINKNKC DIDDLKMAV SFSQFNRRFL NVVRQFSENA GITPAISLDL MTDAELARAI SNMPTSAGQI KLMLENRAMV RRKGFGILIG V YGSSVIYM VQLPIFGVID TPCWIVKAAP SCSEKKGNYA CLLREDQGWY CQNAGSTVYY PNEKDCETRG DHVFCDTAAG IN VAEQSKE CNINISTTNY PCKVSTGRNP ISMVALSPLG ALVACYKGVS CSIGSNRVGI IKQLNKGCSY ITNQDADTVT IDN TVYQLS KVEGEQHVIK GRPVSSSFDP IKFPPDQFNV ALDQVFENIE NSQAWVRKFD EILSSIEKG UniProtKB: Fusion glycoprotein F0 |
-Macromolecule #5: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 5 / Number of copies: 3 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: INSILICO MODEL |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.33 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 453003 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)