+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Voltage gated potassium ion channel Kv1.2 in complex with DTx | |||||||||
 Map data Map data | Kv1.2 DTX bound map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Ion Channel / Toxin bound / Blocker / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationoptic nerve structural organization / Voltage gated Potassium channels / voltage-gated monoatomic ion channel activity involved in regulation of postsynaptic membrane potential / potassium channel complex / paranodal junction / regulation of circadian sleep/wake cycle, non-REM sleep / potassium ion export across plasma membrane / axon initial segment / corpus callosum development / voltage-gated monoatomic ion channel activity involved in regulation of presynaptic membrane potential ...optic nerve structural organization / Voltage gated Potassium channels / voltage-gated monoatomic ion channel activity involved in regulation of postsynaptic membrane potential / potassium channel complex / paranodal junction / regulation of circadian sleep/wake cycle, non-REM sleep / potassium ion export across plasma membrane / axon initial segment / corpus callosum development / voltage-gated monoatomic ion channel activity involved in regulation of presynaptic membrane potential / delayed rectifier potassium channel activity / outward rectifier potassium channel activity / juxtaparanode region of axon / optic nerve development / regulation of dopamine secretion / neuronal cell body membrane / action potential / kinesin binding / lamellipodium membrane / calyx of Held / voltage-gated potassium channel activity / neuronal action potential / potassium channel regulator activity / voltage-gated potassium channel complex / axon terminus / potassium ion transmembrane transport / sensory perception of pain / serine-type endopeptidase inhibitor activity / protein homooligomerization / cerebral cortex development / lamellipodium / presynaptic membrane / toxin activity / perikaryon / postsynaptic membrane / endosome / axon / glutamatergic synapse / dendrite / endoplasmic reticulum membrane / extracellular space / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Komagataella pastoris (fungus) / Komagataella pastoris (fungus) /  Dendroaspis angusticeps (eastern green mamba) / Dendroaspis angusticeps (eastern green mamba) /  | |||||||||
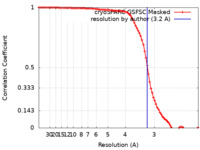
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||
 Authors Authors | Wu Y / Sigworth FJ | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: bioRxiv / Year: 2024 Journal: bioRxiv / Year: 2024Title: Cryo-EM structures of Kv1.2 potassium channels, conducting and non-conducting. Authors: Yangyu Wu / Yangyang Yan / Youshan Yang / Shumin Bian / Alberto Rivetta / Ken Allen / Fred J Sigworth /  Abstract: We present near-atomic-resolution cryo-EM structures of the mammalian voltage-gated potassium channel Kv1.2 in open, C-type inactivated, toxin-blocked and sodium-bound states at 3.2 Å, 2.5 Å, 3.2 ...We present near-atomic-resolution cryo-EM structures of the mammalian voltage-gated potassium channel Kv1.2 in open, C-type inactivated, toxin-blocked and sodium-bound states at 3.2 Å, 2.5 Å, 3.2 Å, and 2.9Å. These structures, all obtained at nominally zero membrane potential in detergent micelles, reveal distinct ion-occupancy patterns in the selectivity filter. The first two structures are very similar to those reported in the related Shaker channel and the much-studied Kv1.2-2.1 chimeric channel. On the other hand, two new structures show unexpected patterns of ion occupancy. First, the toxin α-Dendrotoxin, like Charybdotoxin, is seen to attach to the negatively-charged channel outer mouth, and a lysine residue penetrates into the selectivity filter, with the terminal amine coordinated by carbonyls, partially disrupting the outermost ion-binding site. In the remainder of the filter two densities of bound ions are observed, rather than three as observed with other toxin-blocked Kv channels. Second, a structure of Kv1.2 in Na solution does not show collapse or destabilization of the selectivity filter, but instead shows an intact selectivity filter with ion density in each binding site. We also attempted to image the C-type inactivated Kv1.2 W366F channel in Na solution, but the protein conformation was seen to be highly variable and only a low-resolution structure could be obtained. These findings present new insights into the stability of the selectivity filter and the mechanism of toxin block of this intensively studied, voltage-gated potassium channel. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_43131.map.gz emd_43131.map.gz | 59.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-43131-v30.xml emd-43131-v30.xml emd-43131.xml emd-43131.xml | 14.7 KB 14.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_43131_fsc.xml emd_43131_fsc.xml | 8.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_43131.png emd_43131.png | 75.2 KB | ||
| Filedesc metadata |  emd-43131.cif.gz emd-43131.cif.gz | 5.6 KB | ||
| Others |  emd_43131_half_map_1.map.gz emd_43131_half_map_1.map.gz emd_43131_half_map_2.map.gz emd_43131_half_map_2.map.gz | 59.4 MB 59.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-43131 http://ftp.pdbj.org/pub/emdb/structures/EMD-43131 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43131 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43131 | HTTPS FTP |
-Validation report
| Summary document |  emd_43131_validation.pdf.gz emd_43131_validation.pdf.gz | 1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_43131_full_validation.pdf.gz emd_43131_full_validation.pdf.gz | 1 MB | Display | |
| Data in XML |  emd_43131_validation.xml.gz emd_43131_validation.xml.gz | 16.3 KB | Display | |
| Data in CIF |  emd_43131_validation.cif.gz emd_43131_validation.cif.gz | 21.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43131 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43131 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43131 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43131 | HTTPS FTP |
-Related structure data
| Related structure data |  8vc3MC  8vc4C  8vc6C  8vchC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_43131.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_43131.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Kv1.2 DTX bound map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.068 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Kv1.2 DTX bound half map
| File | emd_43131_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Kv1.2 DTX bound half map | ||||||||||||
| Projections & Slices |
| ||||||||||||
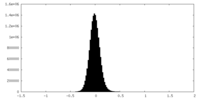
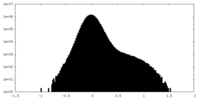
| Density Histograms |
-Half map: Kv1.2 DTX bound half map
| File | emd_43131_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Kv1.2 DTX bound half map | ||||||||||||
| Projections & Slices |
| ||||||||||||
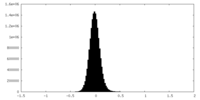
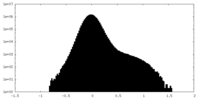
| Density Histograms |
- Sample components
Sample components
-Entire : Voltage gated potassium channel Kv1.2 with DTx bound
| Entire | Name: Voltage gated potassium channel Kv1.2 with DTx bound |
|---|---|
| Components |
|
-Supramolecule #1: Voltage gated potassium channel Kv1.2 with DTx bound
| Supramolecule | Name: Voltage gated potassium channel Kv1.2 with DTx bound / type: cell / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Komagataella pastoris (fungus) / Strain: SMD1168 Komagataella pastoris (fungus) / Strain: SMD1168 |
-Macromolecule #1: Kunitz-type serine protease inhibitor homolog alpha-dendrotoxin
| Macromolecule | Name: Kunitz-type serine protease inhibitor homolog alpha-dendrotoxin type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Dendroaspis angusticeps (eastern green mamba) Dendroaspis angusticeps (eastern green mamba) |
| Molecular weight | Theoretical: 7.015129 KDa |
| Sequence | String: GPRRKLCILH RNPGRCYDKI PAFYYNQKKK QCERFDWSGC GGNSNRFKTI EECRRTCIG UniProtKB: Kunitz-type serine protease inhibitor homolog alpha-dendrotoxin |
-Macromolecule #2: Potassium voltage-gated channel subfamily A member 2
| Macromolecule | Name: Potassium voltage-gated channel subfamily A member 2 / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 60.59157 KDa |
| Recombinant expression | Organism:  Komagataella pastoris (fungus) Komagataella pastoris (fungus) |
| Sequence | String: MSAWSHPQFE KGGGSGGGSG GSAWSHPQFE KLVPRGSMTV ATGDPVDEAA AHPGHPQDTY DPEADHECCE RVVINISGLR FETQLKTLA QFPETLLGDP KKRMRYFDPL RNEYFFDRNR PSFDAILYYY QSGGRLRRPV NVPLDIFSEE IRFYELGEEA M EMFREDEG ...String: MSAWSHPQFE KGGGSGGGSG GSAWSHPQFE KLVPRGSMTV ATGDPVDEAA AHPGHPQDTY DPEADHECCE RVVINISGLR FETQLKTLA QFPETLLGDP KKRMRYFDPL RNEYFFDRNR PSFDAILYYY QSGGRLRRPV NVPLDIFSEE IRFYELGEEA M EMFREDEG YIKEEERPLP ENEFQRQVWL LFEYPESSGP ARIIAIVSVM VILISIVSFC LETLPIFRDE NEDMHGSGVT FH TYSQSTI GYQQSTSFTD PFFIVETLCI IWFSFEFLVR FFACPSKAGF FTNIMNIIDI VAIIPYFITL GTELAEKPED AQQ GQQAMS LAILRVIRLV RVFRIFKLSR HSKGLQILGQ TLKASMRELG LLIFFLFIGV ILFSSAVYFA EADERDSQFP SIPD AFWWA VVSMTTVGYG DMVPTTIGGK IVGSLCAIAG VLTIALPVPV IVSNFNYFYH RETEGEEQAQ YLQVTSCPKI PSSPD LKKS RSASTISKSD YMEIQEGVNN SNEDFREENL KTANCTLANT NYVNITKMLT DV UniProtKB: Potassium voltage-gated channel subfamily A member 2 |
-Macromolecule #3: POTASSIUM ION
| Macromolecule | Name: POTASSIUM ION / type: ligand / ID: 3 / Number of copies: 2 / Formula: K |
|---|---|
| Molecular weight | Theoretical: 39.098 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.1 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)