[English] 日本語
 Yorodumi
Yorodumi- EMDB-42491: CryoEM Structure of Allosterically Switchable De Novo Protein sr3... -
+ Open data
Open data
- Basic information
Basic information
| Entry | 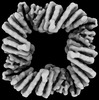 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM Structure of Allosterically Switchable De Novo Protein sr312, in Open State with Effector Peptide | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | de novo / allosterically switchable de novo protein / sr312 / effector peptide / DE NOVO PROTEIN | |||||||||
| Biological species | synthetic construct (others) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.4 Å | |||||||||
 Authors Authors | Weidle C / Skotheim R | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nature Journal: NatureTitle: De novo design of allosterically switchable protein assemblies Authors: Pillai A / Idris A / Philomin A / Weidle C / Skotheim R / Leung PJY / Broerman A / Demakis C / Borst AJ / Praetorius F / Baker D | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_42491.map.gz emd_42491.map.gz | 27.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-42491-v30.xml emd-42491-v30.xml emd-42491.xml emd-42491.xml | 18.9 KB 18.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_42491_fsc.xml emd_42491_fsc.xml | 6.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_42491.png emd_42491.png | 110.1 KB | ||
| Filedesc metadata |  emd-42491.cif.gz emd-42491.cif.gz | 6 KB | ||
| Others |  emd_42491_half_map_1.map.gz emd_42491_half_map_1.map.gz emd_42491_half_map_2.map.gz emd_42491_half_map_2.map.gz | 28.2 MB 28.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-42491 http://ftp.pdbj.org/pub/emdb/structures/EMD-42491 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42491 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42491 | HTTPS FTP |
-Validation report
| Summary document |  emd_42491_validation.pdf.gz emd_42491_validation.pdf.gz | 692.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_42491_full_validation.pdf.gz emd_42491_full_validation.pdf.gz | 692.3 KB | Display | |
| Data in XML |  emd_42491_validation.xml.gz emd_42491_validation.xml.gz | 13.7 KB | Display | |
| Data in CIF |  emd_42491_validation.cif.gz emd_42491_validation.cif.gz | 17.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-42491 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-42491 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-42491 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-42491 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_42491.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_42491.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.2375 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_42491_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_42491_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Complex is comprised of 4 sr312 proteins, each sr312 protein is b...
| Entire | Name: Complex is comprised of 4 sr312 proteins, each sr312 protein is bound to an effector peptide, putting the protein in an open state. Complex is formed with C4 symmetry. |
|---|---|
| Components |
|
-Supramolecule #1: Complex is comprised of 4 sr312 proteins, each sr312 protein is b...
| Supramolecule | Name: Complex is comprised of 4 sr312 proteins, each sr312 protein is bound to an effector peptide, putting the protein in an open state. Complex is formed with C4 symmetry. type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Molecular weight | Theoretical: 206.1 KDa |
-Supramolecule #2: sr312 protein
| Supramolecule | Name: sr312 protein / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
-Supramolecule #3: effector peptide
| Supramolecule | Name: effector peptide / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism: synthetic construct (others) / Synthetically produced: Yes |
-Macromolecule #1: sr312
| Macromolecule | Name: sr312 / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 47.297691 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SGTVTFDITN IDWETAEWIM KHVYLIAKKE GTDVTFSFKE GELQITVKNL HEEAKREIEK WIRAAQLAQD PDAESKAEAR KILNELITE KAEELREKTK DEEVRELARE AARLALESDD IEVQRVVLKA LLAALKSKDE EVIRLLLLAA VLAAAAARSG S PEEKLEIA ...String: SGTVTFDITN IDWETAEWIM KHVYLIAKKE GTDVTFSFKE GELQITVKNL HEEAKREIEK WIRAAQLAQD PDAESKAEAR KILNELITE KAEELREKTK DEEVRELARE AARLALESDD IEVQRVVLKA LLAALKSKDE EVIRLLLLAA VLAAAAARSG S PEEKLEIA KKALELAMKS KDEEVIRLAL LAAVLAARSD DEEVLKKVKE ALEKMERIMD LEDVAREKSG SAEASQAVKE IA DIAEEAL REGLCEVARV ALKRLFKLAK DYPGSDVASL AKKALEKIAE TALRNGCKET AELAKLLLFL LLIIEVVLKM GVR MLTHRG GNAVIVVIEG LHPSQIVQLM QDVIKAAKKL GVTVTITVSG DIVVIMVVVG ASDEEQEEAR RLVQEIARAL QEAK RKGAN EEQLEQLLRE LLERAEREGG SG |
-Macromolecule #2: Effector peptide cs221B
| Macromolecule | Name: Effector peptide cs221B / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 3.091635 KDa |
| Sequence | String: EERKKELAKE VIETAKKLIE KLAKEE |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.0 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
Details: 150 mM NaCl, 40 mM Tris, pH 8.0 | |||||||||
| Grid | Model: C-flat-2/2 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 40 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 25 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 39.0 kPa | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 3795 / Average electron dose: 43.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.8 µm / Nominal defocus min: 0.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



 Z
Z Y
Y X
X