[English] 日本語
 Yorodumi
Yorodumi- EMDB-42114: Cryo-EM structure of the AlbAB cyclodipeptide oxidase enzyme filament -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the AlbAB cyclodipeptide oxidase enzyme filament | |||||||||
 Map data Map data | structure of the AlbAB cyclodipeptide oxidase enzyme filament | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | cyclodipeptide oxidase / cyclic dipeptide oxidase / nitroreductase-like / enzyme filament / flavoenzyme / OXIDOREDUCTASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationalbonoursin synthase / oxidoreductase activity, acting on the CH-CH group of donors, oxygen as acceptor / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Streptomyces noursei ATCC 11455 (bacteria) Streptomyces noursei ATCC 11455 (bacteria) | |||||||||
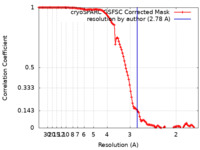
| Method | helical reconstruction / cryo EM / Resolution: 2.78 Å | |||||||||
 Authors Authors | Andreas MP / Giessen TW | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Cyclodipeptide oxidase is an enzyme filament. Authors: Michael P Andreas / Tobias W Giessen /  Abstract: Modified cyclic dipeptides represent a widespread class of secondary metabolites with diverse pharmacological activities, including antibacterial, antifungal, and antitumor. Here, we report the ...Modified cyclic dipeptides represent a widespread class of secondary metabolites with diverse pharmacological activities, including antibacterial, antifungal, and antitumor. Here, we report the structural characterization of the Streptomyces noursei enzyme AlbAB, a cyclodipeptide oxidase (CDO) carrying out α,β-dehydrogenations during the biosynthesis of the antibiotic albonoursin. We show that AlbAB is a megadalton heterooligomeric enzyme filament containing covalently bound flavin mononucleotide cofactors. We highlight that AlbAB filaments consist of alternating dimers of AlbA and AlbB and that enzyme activity is crucially dependent on filament formation. We show that AlbA-AlbB interactions are highly conserved suggesting that other CDO-like enzymes are likely enzyme filaments. As CDOs have been employed in the structural diversification of cyclic dipeptides, our results will be useful for future applications of CDOs in biocatalysis and chemoenzymatic synthesis. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_42114.map.gz emd_42114.map.gz | 167.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-42114-v30.xml emd-42114-v30.xml emd-42114.xml emd-42114.xml | 19.4 KB 19.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_42114_fsc.xml emd_42114_fsc.xml | 11.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_42114.png emd_42114.png | 73.1 KB | ||
| Filedesc metadata |  emd-42114.cif.gz emd-42114.cif.gz | 6.5 KB | ||
| Others |  emd_42114_half_map_1.map.gz emd_42114_half_map_1.map.gz emd_42114_half_map_2.map.gz emd_42114_half_map_2.map.gz | 165.2 MB 165.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-42114 http://ftp.pdbj.org/pub/emdb/structures/EMD-42114 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42114 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42114 | HTTPS FTP |
-Validation report
| Summary document |  emd_42114_validation.pdf.gz emd_42114_validation.pdf.gz | 984.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_42114_full_validation.pdf.gz emd_42114_full_validation.pdf.gz | 983.7 KB | Display | |
| Data in XML |  emd_42114_validation.xml.gz emd_42114_validation.xml.gz | 20.7 KB | Display | |
| Data in CIF |  emd_42114_validation.cif.gz emd_42114_validation.cif.gz | 26.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-42114 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-42114 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-42114 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-42114 | HTTPS FTP |
-Related structure data
| Related structure data |  8uc3MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_42114.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_42114.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | structure of the AlbAB cyclodipeptide oxidase enzyme filament | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.8487 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half Map 1
| File | emd_42114_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half Map 2
| File | emd_42114_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : AlbAB cyclodipeptide oxidase enzyme filament
| Entire | Name: AlbAB cyclodipeptide oxidase enzyme filament |
|---|---|
| Components |
|
-Supramolecule #1: AlbAB cyclodipeptide oxidase enzyme filament
| Supramolecule | Name: AlbAB cyclodipeptide oxidase enzyme filament / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Streptomyces noursei ATCC 11455 (bacteria) Streptomyces noursei ATCC 11455 (bacteria) |
-Macromolecule #1: Albonoursin synthase
| Macromolecule | Name: Albonoursin synthase / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Streptomyces noursei ATCC 11455 (bacteria) Streptomyces noursei ATCC 11455 (bacteria) |
| Molecular weight | Theoretical: 21.071307 KDa |
| Recombinant expression | Organism:  Streptomyces coelicolor A3(2) (bacteria) Streptomyces coelicolor A3(2) (bacteria) |
| Sequence | String: MLAHSSSESP PESLPDAWTV LKTRTAVRNY AKEPVDDALI EQLLEAMLAA PTASNRQAWS FMVVRRPAAV RRLRAFSPGV LGTPAFFVV ACVDRSLTDN LSPKLSQKIY DTSKLCVAMA VENLLLAAHA AGLGGCPVGS FRSDIVTSML GIPEHIEPML V VPIGRPAT ...String: MLAHSSSESP PESLPDAWTV LKTRTAVRNY AKEPVDDALI EQLLEAMLAA PTASNRQAWS FMVVRRPAAV RRLRAFSPGV LGTPAFFVV ACVDRSLTDN LSPKLSQKIY DTSKLCVAMA VENLLLAAHA AGLGGCPVGS FRSDIVTSML GIPEHIEPML V VPIGRPAT ALVPSQRRAK NEVVNYESWG NRAAAPTA UniProtKB: Albonoursin synthase |
-Macromolecule #2: Protein AlbB
| Macromolecule | Name: Protein AlbB / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Streptomyces noursei ATCC 11455 (bacteria) Streptomyces noursei ATCC 11455 (bacteria) |
| Molecular weight | Theoretical: 11.583143 KDa |
| Recombinant expression | Organism:  Streptomyces coelicolor A3(2) (bacteria) Streptomyces coelicolor A3(2) (bacteria) |
| Sequence | String: MNPGETVLPP QLREEIALLA VYLLSSGRGL LEEPADYGIY RCTDGARRAL QLLDEHGGST ARLTAVRERL DEVMFAPMGE DRDMGAILD DLCRQMADAL PEIETP UniProtKB: Protein AlbB |
-Macromolecule #3: FLAVIN MONONUCLEOTIDE
| Macromolecule | Name: FLAVIN MONONUCLEOTIDE / type: ligand / ID: 3 / Number of copies: 2 / Formula: FMN |
|---|---|
| Molecular weight | Theoretical: 456.344 Da |
| Chemical component information |  ChemComp-FMN: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 0.75 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
Details: 20 mM NaCl, 150 mM Tris pH 7.5 | |||||||||
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR Details: Grid was glow discharged for 60 seconds at 5 mA under vacuum | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295 K / Instrument: FEI VITROBOT MARK IV Details: Grid was plunge frozen into liquid ethane using the following parameters: blot force- 5, blot time 2 seconds. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Digitization - Dimensions - Width: 4092 pixel / Digitization - Dimensions - Height: 5760 pixel / Number grids imaged: 1 / Number real images: 3015 / Average exposure time: 2.03063 sec. / Average electron dose: 50.12 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | Chain - Source name: AlphaFold / Chain - Initial model type: in silico model |
|---|---|
| Details | A model of AlbAB was created using AlphaFold2 using MMseqs2 via ColabFold. The model was then fit into the cryoEM density using ChimeraX. The model was then iteratively refined using Coot v 0.9.8.1 and real-space refinement in Phenix v 1.20.1-4487. |
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Overall B value: 107.4 / Target criteria: Cross-correlation coefficient |
| Output model |  PDB-8uc3: |
 Movie
Movie Controller
Controller



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)