[English] 日本語
 Yorodumi
Yorodumi- EMDB-41508: Cryo-EM structure of E3 ubiquitin ligase Doa10 from Saccharomyces... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of E3 ubiquitin ligase Doa10 from Saccharomyces cerevisiae | |||||||||
 Map data Map data | Sharpened map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ubiquitin / ERAD / protein quality control / membrane protein / LIGASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationDoa10p ubiquitin ligase complex / nuclear inner membrane / retrograde protein transport, ER to cytosol / ERAD pathway / RING-type E3 ubiquitin transferase / ubiquitin-protein transferase activity / ubiquitin protein ligase activity / nuclear envelope / protein ubiquitination / endoplasmic reticulum membrane ...Doa10p ubiquitin ligase complex / nuclear inner membrane / retrograde protein transport, ER to cytosol / ERAD pathway / RING-type E3 ubiquitin transferase / ubiquitin-protein transferase activity / ubiquitin protein ligase activity / nuclear envelope / protein ubiquitination / endoplasmic reticulum membrane / endoplasmic reticulum / zinc ion binding / membrane Similarity search - Function | |||||||||
| Biological species |   | |||||||||
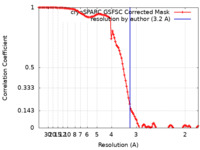
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||
 Authors Authors | Park E / Itskanov SI | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Substrate recognition mechanism of the endoplasmic reticulum-associated ubiquitin ligase Doa10. Authors: Kevin Wu / Samuel Itskanov / Diane L Lynch / Yuanyuan Chen / Aasha Turner / James C Gumbart / Eunyong Park /  Abstract: Doa10 (MARCHF6 in metazoans) is a large polytopic membrane-embedded E3 ubiquitin ligase in the endoplasmic reticulum (ER) that plays an important role in quality control of cytosolic and ER proteins. ...Doa10 (MARCHF6 in metazoans) is a large polytopic membrane-embedded E3 ubiquitin ligase in the endoplasmic reticulum (ER) that plays an important role in quality control of cytosolic and ER proteins. Although Doa10 is highly conserved across eukaryotes, it is not understood how Doa10 recognizes its substrates. Here, we define the substrate recognition mechanism of Doa10 by structural and functional analyses on Saccharomyces cerevisiae Doa10 and its model substrates. Cryo-EM analysis shows that Doa10 has unusual architecture with a large lipid-filled central cavity, and its conserved middle domain forms an additional water-filled lateral tunnel open to the cytosol. Our biochemical data and molecular dynamics simulations suggest that the entrance of the substrate's degron peptide into the lateral tunnel is required for efficient polyubiquitination. The N- and C-terminal membrane domains of Doa10 seem to form fence-like features to restrict polyubiquitination to those proteins that can access the central cavity and lateral tunnel. Our study reveals how extended hydrophobic sequences at the termini of substrate proteins are recognized by Doa10 as a signal for quality control. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_41508.map.gz emd_41508.map.gz | 118.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-41508-v30.xml emd-41508-v30.xml emd-41508.xml emd-41508.xml | 20.4 KB 20.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_41508_fsc.xml emd_41508_fsc.xml | 11.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_41508.png emd_41508.png | 138.2 KB | ||
| Filedesc metadata |  emd-41508.cif.gz emd-41508.cif.gz | 6.6 KB | ||
| Others |  emd_41508_additional_1.map.gz emd_41508_additional_1.map.gz emd_41508_half_map_1.map.gz emd_41508_half_map_1.map.gz emd_41508_half_map_2.map.gz emd_41508_half_map_2.map.gz | 62.3 MB 115.9 MB 115.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-41508 http://ftp.pdbj.org/pub/emdb/structures/EMD-41508 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41508 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41508 | HTTPS FTP |
-Validation report
| Summary document |  emd_41508_validation.pdf.gz emd_41508_validation.pdf.gz | 810.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_41508_full_validation.pdf.gz emd_41508_full_validation.pdf.gz | 810.3 KB | Display | |
| Data in XML |  emd_41508_validation.xml.gz emd_41508_validation.xml.gz | 19 KB | Display | |
| Data in CIF |  emd_41508_validation.cif.gz emd_41508_validation.cif.gz | 24.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41508 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41508 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41508 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41508 | HTTPS FTP |
-Related structure data
| Related structure data |  8tqmMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_41508.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_41508.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.91 Å | ||||||||||||||||||||||||||||||||||||
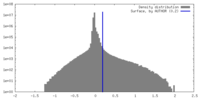
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Unsharpened map
| File | emd_41508_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
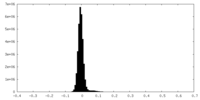
| Density Histograms |
-Half map: #1
| File | emd_41508_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
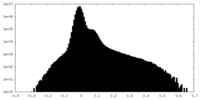
| Density Histograms |
-Half map: #2
| File | emd_41508_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
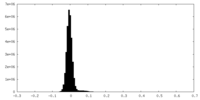
| Density Histograms |
- Sample components
Sample components
-Entire : E3 ubiquitin ligase Doa10
| Entire | Name: E3 ubiquitin ligase Doa10 |
|---|---|
| Components |
|
-Supramolecule #1: E3 ubiquitin ligase Doa10
| Supramolecule | Name: E3 ubiquitin ligase Doa10 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 151 KDa |
-Macromolecule #1: ERAD-associated E3 ubiquitin-protein ligase DOA10
| Macromolecule | Name: ERAD-associated E3 ubiquitin-protein ligase DOA10 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 151.608109 KDa |
| Sequence | String: MDVDSDVNVS RLRDELHKVA NEETDTATFN DDAPSGATCR ICRGEATEDN PLFHPCKCRG SIKYMHESCL LEWVASKNID ISKPGADVK CDICHYPIQF KTIYAENMPE KIPFSLLLSK SILTFFEKAR LALTIGLAAV LYIIGVPLVW NMFGKLYTMM L DGSSPYPG ...String: MDVDSDVNVS RLRDELHKVA NEETDTATFN DDAPSGATCR ICRGEATEDN PLFHPCKCRG SIKYMHESCL LEWVASKNID ISKPGADVK CDICHYPIQF KTIYAENMPE KIPFSLLLSK SILTFFEKAR LALTIGLAAV LYIIGVPLVW NMFGKLYTMM L DGSSPYPG DFLKSLIYGY DQSATPELTT RAIFYQLLQN HSFTSLQFIM IVILHIALYF QYDMIVREDV FSKMVFHKIG PR LSPKDLK SRLKERFPMM DDRMVEYLAR EMRAHDENRQ EQGHDRLNMP AAAADNNNNV INPRNDNVPP QDPNDHRNFE NLR HVDELD HDEATEEHEN NDSDNSLPSG DDSSRILPGS SSDNEEDEEA EGQQQQQQPE EEADYRDHIE PNPIDMWANR RAQN EFDDL IAAQQNAINR PNAPVFIPPP AQNRAGNVDQ DEQDFGAAVG VPPAQANPDD QGQGPLVINL KLKLLNVIAY FIIAV VFTA IYLAISYLFP TFIGFGLLKI YFGIFKVILR GLCHLYYLSG AHIAYNGLTK LVPKVDVAMS WISDHLIHDI IYLYNG YTE NTMKHSIFIR ALPALTTYLT SVSIVCASSN LVSRGYGREN GMSNPTRRLI FQILFALKCT FKVFTLFFIE LAGFPIL AG VMLDFSLFCP ILASNSRMLW VPSICAIWPP FSLFVYWTIG TLYMYWFAKY IGMIRKNIIR PGVLFFIRSP EDPNIKIL H DSLIHPMSIQ LSRLCLSMFI YAIFIVLGFG FHTRIFFPFM LKSNLLSVPE AYKPTSIISW KFNTILLTLY FTKRILESS SYVKPLLERY WKTIFKLCSR KLRLSSFILG KDTPTERGHI VYRNLFYKYI AAKNAEWSNQ ELFTKPKTLE QAEELFGQVR DVHAYFVPD GVLMRVPSSD IVSRNYVQTM FVPVTKDDKL LKPLDLERIK ERNKRAAGEF GYLDEQNTEY DQYYIVYVPP D FRLRYMTL LGLVWLFASI LMLGVTFISQ ALINFVCSFG FLPVVKLLLG ERNKVYVAWK ELSDISYSYL NIYYVCVGSV CL SKIAKDI LHFTEGQNTL DEHAVDENEV EEVEHDIPER DINNAPVNNI NNVEEGQGIF MAIFNSIFDS MLVKYNLMVF IAI MIAVIR TMVSWVVLTD GILACYNYLT IRVFGNSSYT IGNSKWFKYD ESLLFVVWII SSMVNFGTGY KSLKLFFRNR NTSK LNFLK TMALELFKQG FLHMVIYVLP IIILSLVFLR DVSTKQIIDI SHGSRSFTLS LNESFPTWTR MQDIYFGLLI ALESF TFFF QATVLFIQWF KSTVQNVKDE VYTKGRALEN LPDES UniProtKB: ERAD-associated E3 ubiquitin-protein ligase DOA10 |
-Macromolecule #2: 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE
| Macromolecule | Name: 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE / type: ligand / ID: 2 / Number of copies: 4 / Formula: PC1 |
|---|---|
| Molecular weight | Theoretical: 790.145 Da |
| Chemical component information |  ChemComp-PC1: |
-Macromolecule #3: TRISTEAROYLGLYCEROL
| Macromolecule | Name: TRISTEAROYLGLYCEROL / type: ligand / ID: 3 / Number of copies: 1 / Formula: TGL |
|---|---|
| Molecular weight | Theoretical: 891.48 Da |
| Chemical component information |  ChemComp-TGL: |
-Macromolecule #4: CHOLESTEROL HEMISUCCINATE
| Macromolecule | Name: CHOLESTEROL HEMISUCCINATE / type: ligand / ID: 4 / Number of copies: 1 / Formula: Y01 |
|---|---|
| Molecular weight | Theoretical: 486.726 Da |
| Chemical component information |  ChemComp-Y01: |
-Macromolecule #5: ERGOSTEROL
| Macromolecule | Name: ERGOSTEROL / type: ligand / ID: 5 / Number of copies: 1 / Formula: ERG |
|---|---|
| Molecular weight | Theoretical: 396.648 Da |
| Chemical component information |  ChemComp-ERG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 5.5 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
| Details | 20 mM Tris-HCl, pH 7.5, 100 mM NaCl, 1 mM EDTA, 2 mM DTT, 0.02% GDN |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.4 µm / Nominal defocus min: 0.7000000000000001 µm / Nominal magnification: 64000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | Chain - Source name: Other / Chain - Initial model type: other / Details: de novo |
|---|---|
| Refinement | Protocol: AB INITIO MODEL |
| Output model |  PDB-8tqm: |
 Movie
Movie Controller
Controller



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)