+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Cryo-EM structure of DDB1dB:CRBN:Pomalidomide:SD40 | |||||||||
 マップデータ マップデータ | main map | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | ubiquitin / CRBN / directed evolution / Zinc finger / IMiD / Molecular Glue / TRANSFERASE | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報negative regulation of monoatomic ion transmembrane transport / lymphocyte differentiation / positive regulation by virus of viral protein levels in host cell / spindle assembly involved in female meiosis / epigenetic programming in the zygotic pronuclei / Cul4-RING E3 ubiquitin ligase complex / UV-damage excision repair / biological process involved in interaction with symbiont / regulation of mitotic cell cycle phase transition / NOTCH3 Intracellular Domain Regulates Transcription ...negative regulation of monoatomic ion transmembrane transport / lymphocyte differentiation / positive regulation by virus of viral protein levels in host cell / spindle assembly involved in female meiosis / epigenetic programming in the zygotic pronuclei / Cul4-RING E3 ubiquitin ligase complex / UV-damage excision repair / biological process involved in interaction with symbiont / regulation of mitotic cell cycle phase transition / NOTCH3 Intracellular Domain Regulates Transcription / WD40-repeat domain binding / Cul4A-RING E3 ubiquitin ligase complex / Cul4B-RING E3 ubiquitin ligase complex / ubiquitin ligase complex scaffold activity / detection of maltose stimulus / maltose transport complex / negative regulation of reproductive process / negative regulation of developmental process / mesoderm development / locomotory exploration behavior / maltose binding / carbohydrate transport / maltose transport / viral release from host cell / maltodextrin transmembrane transport / cullin family protein binding / positive regulation of Wnt signaling pathway / ectopic germ cell programmed cell death / carbohydrate transmembrane transporter activity / ATP-binding cassette (ABC) transporter complex, substrate-binding subunit-containing / negative regulation of protein-containing complex assembly / proteasomal protein catabolic process / positive regulation of viral genome replication / pericentric heterochromatin / positive regulation of gluconeogenesis / ATP-binding cassette (ABC) transporter complex / cell chemotaxis / erythrocyte differentiation / nucleotide-excision repair / Recognition of DNA damage by PCNA-containing replication complex / positive regulation of protein-containing complex assembly / DNA Damage Recognition in GG-NER / regulation of circadian rhythm / Transcription-Coupled Nucleotide Excision Repair (TC-NER) / Formation of TC-NER Pre-Incision Complex / Dual Incision in GG-NER / Wnt signaling pathway / Formation of Incision Complex in GG-NER / Dual incision in TC-NER / Gap-filling DNA repair synthesis and ligation in TC-NER / positive regulation of protein catabolic process / cellular response to UV / rhythmic process / chromatin organization / site of double-strand break / Neddylation / protein-macromolecule adaptor activity / outer membrane-bounded periplasmic space / ubiquitin-dependent protein catabolic process / proteasome-mediated ubiquitin-dependent protein catabolic process / transmembrane transporter binding / Potential therapeutics for SARS / damaged DNA binding / chromosome, telomeric region / periplasmic space / protein ubiquitination / DNA-binding transcription factor activity / RNA polymerase II cis-regulatory region sequence-specific DNA binding / protein domain specific binding / DNA repair / negative regulation of DNA-templated transcription / DNA damage response / protein-containing complex binding / negative regulation of apoptotic process / regulation of transcription by RNA polymerase II / nucleolus / apoptotic process / perinuclear region of cytoplasm / protein-containing complex / DNA binding / extracellular space / extracellular exosome / nucleoplasm / identical protein binding / membrane / nucleus / metal ion binding / cytosol / cytoplasm 類似検索 - 分子機能 | |||||||||
| 生物種 |  Homo sapiens (ヒト) Homo sapiens (ヒト) | |||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 3.3 Å | |||||||||
 データ登録者 データ登録者 | Roy Burman SS / Hunkeler M / Fischer ES | |||||||||
| 資金援助 |  米国, 2件 米国, 2件
| |||||||||
 引用 引用 |  ジャーナル: Science / 年: 2024 ジャーナル: Science / 年: 2024タイトル: Continuous evolution of compact protein degradation tags regulated by selective molecular glues. 著者: Jaron A M Mercer / Stephan J DeCarlo / Shourya S Roy Burman / Vedagopuram Sreekanth / Andrew T Nelson / Moritz Hunkeler / Peter J Chen / Katherine A Donovan / Praveen Kokkonda / Praveen K ...著者: Jaron A M Mercer / Stephan J DeCarlo / Shourya S Roy Burman / Vedagopuram Sreekanth / Andrew T Nelson / Moritz Hunkeler / Peter J Chen / Katherine A Donovan / Praveen Kokkonda / Praveen K Tiwari / Veronika M Shoba / Arghya Deb / Amit Choudhary / Eric S Fischer / David R Liu /  要旨: Conditional protein degradation tags (degrons) are usually >100 amino acids long or are triggered by small molecules with substantial off-target effects, thwarting their use as specific modulators of ...Conditional protein degradation tags (degrons) are usually >100 amino acids long or are triggered by small molecules with substantial off-target effects, thwarting their use as specific modulators of endogenous protein levels. We developed a phage-assisted continuous evolution platform for molecular glue complexes (MG-PACE) and evolved a 36-amino acid zinc finger (ZF) degron (SD40) that binds the ubiquitin ligase substrate receptor cereblon in complex with PT-179, an orthogonal thalidomide derivative. Endogenous proteins tagged in-frame with SD40 using prime editing are degraded by otherwise inert PT-179. Cryo-electron microscopy structures of SD40 in complex with ligand-bound cereblon revealed mechanistic insights into the molecular basis of SD40's activity and specificity. Our efforts establish a system for continuous evolution of molecular glue complexes and provide ZF tags that overcome shortcomings associated with existing degrons. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_41423.map.gz emd_41423.map.gz | 40.5 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-41423-v30.xml emd-41423-v30.xml emd-41423.xml emd-41423.xml | 41.2 KB 41.2 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_41423_fsc.xml emd_41423_fsc.xml | 7.4 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_41423.png emd_41423.png | 90.3 KB | ||
| マスクデータ |  emd_41423_msk_1.map emd_41423_msk_1.map | 42.9 MB |  マスクマップ マスクマップ | |
| Filedesc metadata |  emd-41423.cif.gz emd-41423.cif.gz | 8.7 KB | ||
| その他 |  emd_41423_additional_1.map.gz emd_41423_additional_1.map.gz emd_41423_additional_2.map.gz emd_41423_additional_2.map.gz emd_41423_additional_3.map.gz emd_41423_additional_3.map.gz emd_41423_additional_4.map.gz emd_41423_additional_4.map.gz emd_41423_additional_5.map.gz emd_41423_additional_5.map.gz emd_41423_additional_6.map.gz emd_41423_additional_6.map.gz emd_41423_additional_7.map.gz emd_41423_additional_7.map.gz emd_41423_half_map_1.map.gz emd_41423_half_map_1.map.gz emd_41423_half_map_2.map.gz emd_41423_half_map_2.map.gz | 1.1 MB 39.8 MB 39.8 MB 38.4 MB 21.8 MB 38.5 MB 40.5 MB 39.8 MB 39.8 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-41423 http://ftp.pdbj.org/pub/emdb/structures/EMD-41423 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41423 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41423 | HTTPS FTP |
-検証レポート
| 文書・要旨 |  emd_41423_validation.pdf.gz emd_41423_validation.pdf.gz | 1.1 MB | 表示 |  EMDB検証レポート EMDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  emd_41423_full_validation.pdf.gz emd_41423_full_validation.pdf.gz | 1.1 MB | 表示 | |
| XML形式データ |  emd_41423_validation.xml.gz emd_41423_validation.xml.gz | 15.1 KB | 表示 | |
| CIF形式データ |  emd_41423_validation.cif.gz emd_41423_validation.cif.gz | 19 KB | 表示 | |
| アーカイブディレクトリ |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41423 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41423 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41423 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41423 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  8tnpMC  8tnqC  8tnrC C: 同じ文献を引用 ( M: このマップから作成された原子モデル |
|---|---|
| 類似構造データ | 類似検索 - 機能・相同性  F&H 検索 F&H 検索 |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_41423.map.gz / 形式: CCP4 / 大きさ: 42.9 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_41423.map.gz / 形式: CCP4 / 大きさ: 42.9 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | main map | ||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.277 Å | ||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
+マスク #1
+追加マップ: main map, local resolution-filtered
+追加マップ: from local refinement with CRBN mask, half map A
+追加マップ: from local refinement with CRBN mask, half map B
+追加マップ: map from local refinement, post-processed with deepEMhancer
+追加マップ: unsharpened main map
+追加マップ: main map, post-processed by deepEMhancer
+追加マップ: from local refinement with CRBN mask
+ハーフマップ: half map A
+ハーフマップ: half map B
- 試料の構成要素
試料の構成要素
-全体 : Ternary Complex of DDB1dB:CRBN:pomalidomide with SD40
| 全体 | 名称: Ternary Complex of DDB1dB:CRBN:pomalidomide with SD40 |
|---|---|
| 要素 |
|
-超分子 #1: Ternary Complex of DDB1dB:CRBN:pomalidomide with SD40
| 超分子 | 名称: Ternary Complex of DDB1dB:CRBN:pomalidomide with SD40 タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: #1-#3 |
|---|---|
| 分子量 | 理論値: 201 KDa |
-分子 #1: Protein cereblon
| 分子 | 名称: Protein cereblon / タイプ: protein_or_peptide / ID: 1 / コピー数: 1 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 55.144594 KDa |
| 組換発現 | 生物種:  Trichoplusia ni (イラクサキンウワバ) Trichoplusia ni (イラクサキンウワバ) |
| 配列 | 文字列: MDYKDDDDKS AVDENLYFQG GGRGGSAHIV MVDAYKPTKG GSGMAGEGDQ QDAAHNMGNH LPLLPAESEE EDEMEVEDQD SKEAKKPNI INFDTSLPTS HTYLGADMEE FHGRTLHDDD SCQVIPVLPQ VMMILIPGQT LPLQLFHPQE VSMVRNLIQK D RTFAVLAY ...文字列: MDYKDDDDKS AVDENLYFQG GGRGGSAHIV MVDAYKPTKG GSGMAGEGDQ QDAAHNMGNH LPLLPAESEE EDEMEVEDQD SKEAKKPNI INFDTSLPTS HTYLGADMEE FHGRTLHDDD SCQVIPVLPQ VMMILIPGQT LPLQLFHPQE VSMVRNLIQK D RTFAVLAY SNVQEREAQF GTTAEIYAYR EEQDFGIEIV KVKAIGRQRF KVLELRTQSD GIQQAKVQIL PECVLPSTMS AV QLESLNK CQIFPSKPVS REDQCSYKWW QKYQKRKFHC ANLTSWPRWL YSLYDAETLM DRIKKQLREW DENLKDDSLP SNP IDFSYR VAACLPIDDV LRIQLLKIGS AIQRLRCELD IMNKCTSLCC KQCQETEITT KNEIFSLSLC GPMAAYVNPH GYVH ETLTV YKACNLNLIG RPSTEHSWFP GYAWTVAQCK ICASHIGWKF TATKKDMSPQ KFWGLTRSAL LPTIPDTEDE ISPDK VILC L UniProtKB: Protein cereblon |
-分子 #2: DNA damage-binding protein 1
| 分子 | 名称: DNA damage-binding protein 1 / タイプ: protein_or_peptide / ID: 2 / コピー数: 1 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 96.033945 KDa |
| 組換発現 | 生物種:  Trichoplusia ni (イラクサキンウワバ) Trichoplusia ni (イラクサキンウワバ) |
| 配列 | 文字列: MGSSHHHHHH SAVDENLYFQ GGGRMSYNYV VTAQKPTAVN GCVTGHFTSA EDLNLLIAKN TRLEIYVVTA EGLRPVKEVG MYGKIAVME LFRPKGESKD LLFILTAKYN ACILEYKQSG ESIDIITRAH GNVQDRIGRP SETGIIGIID PECRMIGLRL Y DGLFKVIP ...文字列: MGSSHHHHHH SAVDENLYFQ GGGRMSYNYV VTAQKPTAVN GCVTGHFTSA EDLNLLIAKN TRLEIYVVTA EGLRPVKEVG MYGKIAVME LFRPKGESKD LLFILTAKYN ACILEYKQSG ESIDIITRAH GNVQDRIGRP SETGIIGIID PECRMIGLRL Y DGLFKVIP LDRDNKELKA FNIRLEELHV IDVKFLYGCQ APTICFVYQD PQGRHVKTYE VSLREKEFNK GPWKQENVEA EA SMVIAVP EPFGGAIIIG QESITYHNGD KYLAIAPPII KQSTIVCHNR VDPNGSRYLL GDMEGRLFML LLEKEEQMDG TVT LKDLRV ELLGETSIAE CLTYLDNGVV FVGSRLGDSQ LVKLNVDSNE QGSYVVAMET FTNLGPIVDM CVVDLERQGQ GQLV TCSGA FKEGSLRIIR NGIGGNGNSG EIQKLHIRTV PLYESPRKIC YQEVSQCFGV LSSRIEVQDT SGGTTALRPS ASTQA LSSS VSSSKLFSSS TAPHETSFGE EVEVHNLLII DQHTFEVLHA HQFLQNEYAL SLVSCKLGKD PNTYFIVGTA MVYPEE AEP KQGRIVVFQY SDGKLQTVAE KEVKGAVYSM VEFNGKLLAS INSTVRLYEW TTEKELRTEC NHYNNIMALY LKTKGDF IL VGDLMRSVLL LAYKPMEGNF EEIARDFNPN WMSAVEILDD DNFLGAENAF NLFVCQKDSA ATTDEERQHL QEVGLFHL G EFVNVFCHGS LVMQNLGETS TPTQGSVLFG TVNGMIGLVT SLSESWYNLL LDMQNRLNKV IKSVGKIEHS FWRSFHTER KTEPATGFID GDLIESFLDI SRPKMQEVVA NLQYDDGSGM KREATADDLI KVVEELTRIH UniProtKB: DNA damage-binding protein 1, DNA damage-binding protein 1 |
-分子 #3: Maltose/maltodextrin-binding periplasmic protein,SD40
| 分子 | 名称: Maltose/maltodextrin-binding periplasmic protein,SD40 タイプ: protein_or_peptide / ID: 3 詳細: IZKF1/ZFP91 fusion construct that was further engineered to enhance binding of cereblon/DDB1 in the presence of IMiD derivatives コピー数: 1 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 50.466012 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MGLNDIFEAQ KIEWHEGSSH HHHHHGSSKI EEGKLVIWIN GDKGYNGLAE VGKKFEKDTG IKVTVEHPDK LEEKFPQVAA TGDGPDIIF WAHDRFGGYA QSGLLAEITP DKAFQDKLYP FTWDAVRYNG KLIAYPIAVE ALSLIYNKDL LPNPPKTWEE I PALDKELK ...文字列: MGLNDIFEAQ KIEWHEGSSH HHHHHGSSKI EEGKLVIWIN GDKGYNGLAE VGKKFEKDTG IKVTVEHPDK LEEKFPQVAA TGDGPDIIF WAHDRFGGYA QSGLLAEITP DKAFQDKLYP FTWDAVRYNG KLIAYPIAVE ALSLIYNKDL LPNPPKTWEE I PALDKELK AKGKSALMFN LQEPYFTWPL IAADGGYAFK YENGKYDIKD VGVDNAGAKA GLTFLVDLIK NKHMNADTDY SI AEAAFNK GETAMTINGP WAWSNIDTSK VNYGVTVLPT FKGQPSKPFV GVLSAGINAA SPNKELAKEF LENYLLTDEG LEA VNKDKP LGAVALKSYE EELAKDPRIA ATMENAQKGE IMPNIPQMSA FWYAVRTAVI NAASGRQTVD EALKDAQTRI TKLE VLFQG PDYKDDDDKS GGGGLLLFCP ICGFTCRQKG NLLRHINLHT GEKLFKYHLY UniProtKB: Maltose/maltodextrin-binding periplasmic protein, DNA-binding protein Ikaros |
-分子 #4: ZINC ION
| 分子 | 名称: ZINC ION / タイプ: ligand / ID: 4 / コピー数: 2 / 式: ZN |
|---|---|
| 分子量 | 理論値: 65.409 Da |
-分子 #5: S-Pomalidomide
| 分子 | 名称: S-Pomalidomide / タイプ: ligand / ID: 5 / コピー数: 1 / 式: Y70 |
|---|---|
| 分子量 | 理論値: 273.244 Da |
| Chemical component information | 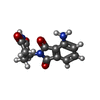 ChemComp-Y70: |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 濃度 | 0.2625 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 緩衝液 | pH: 7 構成要素:
詳細: 20 mM HEPES/NaOH pH 7.0, 150 mM NaCl, and 3 mM TCEP. DMSO concentrations were kept below 2% (v/v) | ||||||||||||
| グリッド | モデル: Quantifoil R1.2/1.3 / 材質: GOLD / メッシュ: 300 / 支持フィルム - 材質: GOLD / 支持フィルム - トポロジー: HOLEY ARRAY / 支持フィルム - Film thickness: 50 / 前処理 - タイプ: GLOW DISCHARGE / 前処理 - 時間: 120 sec. / 前処理 - 雰囲気: AIR / 前処理 - 気圧: 0.039 kPa 詳細: Grids (Quantifoil UltrAuFoil R 1.2/1.3) were glow discharged in PELCO easiGlow (20 mA, 120s, 39 Pa) | ||||||||||||
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 90 % / チャンバー内温度: 283 K / 装置: LEICA EM GP 詳細: Leica EM-GP plunge freezer with chamber conditions of 10 C and 90% relative humidity. Grids were first pre-incubated with 4 uL of 10 uM CRBN-agnostic IKZF1_140-196_Q146A,G151N for 1 minute ...詳細: Leica EM-GP plunge freezer with chamber conditions of 10 C and 90% relative humidity. Grids were first pre-incubated with 4 uL of 10 uM CRBN-agnostic IKZF1_140-196_Q146A,G151N for 1 minute and then blotted from behind for 4 s. Immediately, 4 uL of mixture 1 diluted 10-fold--with the dilution buffer during the 1-minute incubation time--was applied to the grids before blotting for 4 s and plunging into liquid ethane at -181 C.. | ||||||||||||
| 詳細 | DDB1dB_CRBN, pomalidomide, and SD40 were mixed and incubated on ice for 1 hour at final concentration of 10.5, 105, and 21 uM, respectively. Then diluted 10-fold before blotting. |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | TFS TALOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K3 (6k x 4k) / 撮影したグリッド数: 1 / 実像数: 2524 / 平均露光時間: 4.993 sec. / 平均電子線量: 53.8 e/Å2 詳細: Movies (50 frames) collected using beam-shift with 9 holes per stage position (3x3 pattern) |
| 電子線 | 加速電圧: 200 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | C2レンズ絞り径: 50.0 µm / 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / Cs: 2.7 mm / 最大 デフォーカス(公称値): 2.0 µm / 最小 デフォーカス(公称値): 0.8 µm / 倍率(公称値): 36000 |
| 試料ステージ | 試料ホルダーモデル: FEI TITAN KRIOS AUTOGRID HOLDER ホルダー冷却材: NITROGEN |
+ 画像解析
画像解析
-原子モデル構築 1
| 初期モデル |
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 詳細 | Phenix real-space refinement without rigid body | ||||||||
| 精密化 | 空間: REAL / プロトコル: OTHER / 温度因子: 102 / 当てはまり具合の基準: CC | ||||||||
| 得られたモデル |  PDB-8tnp: |
 ムービー
ムービー コントローラー
コントローラー





















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)